CCR5 and the Δ32 Mutation: From Natural HIV Resistance to Cutting-Edge Cure Strategies
This article provides a comprehensive analysis of the CCR5 co-receptor and its pivotal role in HIV-1 entry.
CCR5 and the Δ32 Mutation: From Natural HIV Resistance to Cutting-Edge Cure Strategies
Abstract
This article provides a comprehensive analysis of the CCR5 co-receptor and its pivotal role in HIV-1 entry. It explores the foundational biology of CCR5, the protective mechanism of the natural CCR5-Δ32 mutation, and its population genetics. The review systematically evaluates current methodological advances, including CCR5-targeted gene editing technologies (ZFNs, TALENs, CRISPR/Cas9) and their application in therapeutic and cure strategies, such as stem cell transplantation. It further addresses key challenges in the field, including viral tropism switching, off-target effects of gene editing, and the pleiotropic health impacts of CCR5 disruption. Finally, it offers a comparative validation of these approaches against traditional antiretroviral therapy, synthesizing efficacy and safety data from clinical and preclinical studies to present a roadmap for the future of HIV treatment and cure research aimed at scientists, researchers, and drug development professionals.
The Gateway to Infection: Unraveling CCR5 Biology and the Protective Δ32 Mutation
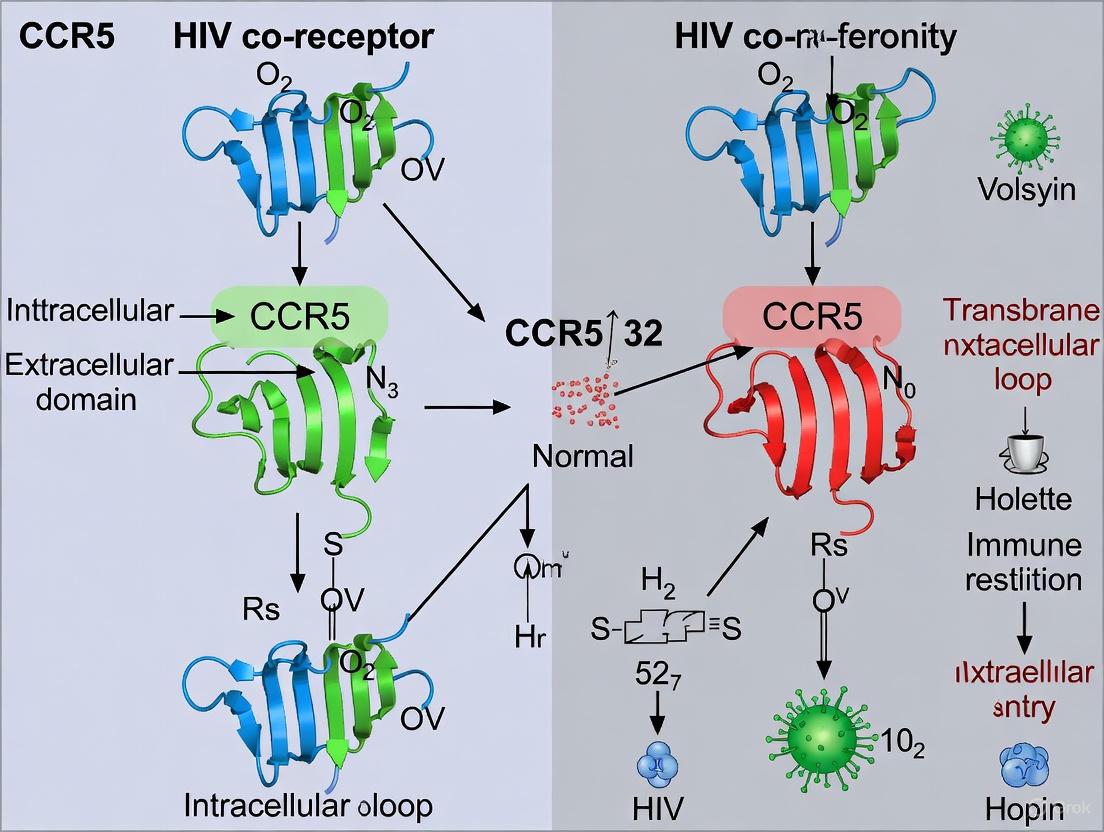
C-C chemokine receptor 5 (CCR5) serves as the principal coreceptor for human immunodeficiency virus type 1 (HIV-1) entry, representing a critical determinant of viral transmission and pathogenesis. This seven-transmembrane G protein-coupled receptor (GPCR), expressed predominantly on immune cells such as T lymphocytes and macrophages, facilitates HIV-1 fusion through specific interactions with the viral envelope glycoprotein gp120 following its initial attachment to CD4. The discovery of the CCR5Δ32 mutation and its association with natural resistance to HIV-1 infection has catalyzed extensive research into CCR5-targeted therapeutic strategies. This whitepaper provides a comprehensive technical analysis of CCR5 structure-function relationships, detailed experimental methodologies for investigating coreceptor activity, and current therapeutic approaches including small molecule antagonists and gene editing technologies, framed within the context of advancing a functional cure for HIV-1 infection.
The identification of CCR5 as an essential coreceptor for HIV-1 entry marked a pivotal advancement in AIDS research, explaining viral tropism and providing novel therapeutic targets. HIV-1 infection requires sequential interactions between the viral envelope glycoprotein gp120, the primary receptor CD4, and a coreceptor—predominantly CCR5 or CXCR4. Macrophage-tropic (R5) strains, which utilize CCR5, constitute the predominantly transmitted variants and are established during early infection [1] [2]. The critical role of CCR5 in HIV-1 pathogenesis is empirically demonstrated by the natural resistance observed in individuals carrying the homozygous CCR5-Δ32 mutation, which truncates and inactivates the receptor [3] [4]. This protective effect, famously documented in the "Berlin" and "London" HIV cure cases following CCR5-Δ32 hematopoietic stem cell transplantation, provides a compelling rationale for CCR5-targeted therapeutic interventions [3].
Beyond its role in HIV-1 entry, CCR5 functions as a typical chemokine receptor, binding inflammatory ligands including MIP-1α (CCL3), MIP-1β (CCL4), and RANTES (CCL5) [5] [4]. These endogenous chemokines can competitively inhibit HIV-1 entry in vitro, suggesting natural mechanisms of viral suppression [4]. CCR5 regulates leukocyte chemotaxis to inflammatory sites and is implicated in various inflammatory diseases, though its absence in Δ32 carriers presents no overt pathological phenotype, making it an attractive and potentially safe therapeutic target [5] [2].
Structural Organization of CCR5
Canonical GPCR Architecture
CCR5 belongs to the Class A rhodopsin-like GPCR family, characterized by a conserved structural framework of seven transmembrane (7TM) α-helices connected by three extracellular loops (ECL1-3) and three intracellular loops (ICL1-3) [4]. The receptor features an extracellular N-terminus important for ligand binding and an intracellular C-terminus involved in signaling regulation. Two conserved disulfide bridges stabilize the extracellular domain: one between Cys101 in helix III and Cys178 in ECL2, and another connecting Cys20 in the N-terminus to Cys269 in helix VII [4]. This structural arrangement creates a ligand-binding pocket capable of accommodating both chemokines and viral envelope proteins.
Table 1: Key Structural Elements of CCR5 and Their Functional Roles
| Structural Element | Composition | Functional Role in HIV-1 Entry |
|---|---|---|
| N-terminal domain | Tyrosine-rich region (Tyr10, Tyr14, Tyr15) | Critical for gp120 binding; sulfation of tyrosine residues enhances HIV-1 affinity [1] |
| Extracellular loops | ECL2 (β-hairpin structure) | Forms principal contact surface for gp120; determines coreceptor specificity [2] |
| Transmembrane helices | 7 α-helices bundle | Creates deep binding pocket for small molecule inhibitors (e.g., maraviroc) [4] |
| Intracellular domains | C-terminus and ICLs | Mediates G-protein coupling and signal transduction; regulates receptor internalization [5] |
Conformational Dynamics and Plasticity
CCR5 exhibits significant structural plasticity, existing in multiple conformational states that influence its functions [1]. Emerging evidence reveals that CCR5 can adopt various conformations that are not equivalent in terms of trafficking, signaling, and coreceptor activity [2]. Monoclonal antibodies directed at different CCR5 epitopes demonstrate varying capacities to inhibit HIV-1 infection, suggesting the virus preferentially utilizes specific CCR5 conformations for entry [2]. This conformational diversity stems from several factors:
- Post-translational modifications: Sulfation of N-terminal tyrosine residues (3, 10, 14, 15) enhances binding to gp120/CD4 complexes [1]. Additionally, palmitoylation of cysteine residues in the C-terminal tail regulates receptor trafficking and signaling efficiency [2].
- Oligomerization: CCR5 forms homodimers and heterodimers with other receptors, including CD4, which can influence HIV-1 entry efficiency by facilitating the formation of entry complexes at the plasma membrane [2].
- Membrane composition: The lipid microenvironment influences CCR5 conformation and function, with cholesterol-rich lipid rafts particularly facilitating HIV-1 entry [1].
Molecular Mechanism of HIV-1 Entry via CCR5
Sequential Binding Process
HIV-1 entry through CCR5 follows a meticulously orchestrated sequence of molecular interactions that ultimately lead to viral-cell membrane fusion:
Primary CD4 attachment: The viral gp120 glycoprotein initially engages the CD4 receptor on the target cell surface, inducing conformational changes in gp120 that expose previously cryptic epitopes, including the bridging sheet and V3 loop [6] [4].
Coreceptor engagement: The structurally remodeled gp120 then interacts with CCR5, primarily through contacts between the V3 loop and the CCR5 N-terminus, with additional interactions involving the bridging sheet and ECL2 [4]. The tyrosine-sulfated N-terminus of CCR5 represents an essential determinant for gp120 binding [4].
Fusion activation: CCR5 binding triggers additional conformational changes in the envelope glycoprotein complex, leading to the exposure and insertion of the gp41 fusion peptide into the target cell membrane [6].
Membrane fusion: gp41 refolds into a stable six-helix bundle structure, bringing the viral and cellular membranes into close proximity and catalyzing their fusion, resulting in viral capsid entry into the cytoplasm [6].
Diagram Title: Sequential Process of HIV-1 Entry via CCR5
Key Interaction Domains
The molecular interface between gp120 and CCR5 involves several structurally defined regions:
- CCR5 N-terminus: The sulfated tyrosine residues at positions 10, 14, and 15 provide critical electrostatic interactions with gp120 [1] [4].
- V3 loop of gp120: This hypervariable region determines coreceptor specificity, with basic residues facilitating CCR5 engagement [1] [4].
- ECL2 of CCR5: The second extracellular loop forms a β-hairpin structure that interacts with the gp120 bridging sheet [2] [4].
- Transmembrane pocket: Small molecule antagonists like maraviroc bind deep within the TM helix bundle, stabilizing CCR5 in an inactive conformation incompatible with gp120 binding [4].
The CCR5-Δ32 Mutation: Natural Resistance Mechanism
The CCR5-Δ32 mutation represents a 32-base pair deletion in the CCR5 gene open reading frame, resulting in a frameshift and premature termination codon that produces a truncated, non-functional receptor retained intracellularly [4]. This mutation confers natural resistance to R5-tropic HIV-1 infection through distinct mechanisms depending on genotype:
- Δ32 homozygotes (approximately 1% of Caucasian populations): Complete absence of functional CCR5 on cell surfaces provides near-complete protection against R5-tropic HIV-1 transmission [4].
- Δ32 heterozygotes (approximately 10-15% of Caucasian populations): Reduced CCR5 cell surface expression results in slowed disease progression and lower viral loads following infection [2].
The Δ32 mutation exhibits a geographical gradient in allele frequency, with highest prevalence in Northern European populations (up to 16%) and decreasing toward Southern Europe and Asia, suggesting historical selective pressure, possibly from pathogens like plague or smallpox [2]. The protective effect of CCR5 deficiency against HIV-1 without apparent severe immunodeficiency has made CCR5 an ideal target for therapeutic intervention.
Quantitative Data on CCR5 Expression and Function
Table 2: Quantitative Parameters of CCR5 Expression and HIV-1 Entry
| Parameter | Value/Range | Biological Significance | Reference |
|---|---|---|---|
| Protein size | 40-45 kDa (352 amino acids) | Standard for GPCRs; varies with glycosylation | [4] |
| CCR5 copies/cell on primary CD4+ T cells | 1,000 - 50,000 molecules | Higher expression correlates with increased susceptibility to HIV-1 infection | [2] |
| CCR5-Δ32 allele frequency (Northern Europe) | 10-16% | Population-specific protective effect against HIV-1 | [2] |
| Δ32 heterozygote CCR5 surface expression | ~30-50% reduction | Slower disease progression in HIV+ individuals | [2] |
| Contrast ratio for large text | ≥4.5:1 | Accessibility standard for scientific diagrams | [7] [8] |
| Contrast ratio for standard text | ≥7:1 | Accessibility standard for scientific text | [7] [8] |
Experimental Protocols for CCR5 Research
Coreceptor Function Assay
Purpose: To quantitatively evaluate CCR5-dependent HIV-1 entry into target cells.
Methodology:
- Cell preparation: Use CCR5-expressing cell lines (e.g., PM1, U87.CD4.CCR5) or primary CD4+ T cells activated with PHA/IL-2.
- Virus inoculation: Incubate cells with R5-tropic HIV-1 strains (e.g., BaL, JR-FL) at specified multiplicity of infection (MOI: 0.01-0.1) for 2-4 hours at 37°C.
- Inhibition controls: Include:
- CCR5 antagonists (maraviroc, 1-10 μM)
- Neutralizing anti-CCR5 antibodies (e.g., 2D7, 10 μg/mL)
- Natural chemokine ligands (RANTES, 100 nM)
- Infection readout:
- Intracellular p24 staining: Fix and permeabilize cells at 48 hours post-infection; stain with anti-p24-FITC; analyze by flow cytometry.
- Luciferase reporter assay: Use pseudotyped viruses with luciferase reporter gene; measure luminescence at 72 hours post-infection.
- Data analysis: Calculate percentage infection inhibition relative to untreated controls using formula:
% Inhibition = [1 - (Experimental/Control)] × 100[1] [2].
CCR5 Gene Editing Validation Protocol
Purpose: To verify efficient CCR5 disruption using CRISPR/Cas9 or other gene editing technologies.
Methodology:
- Guide RNA design: Target CCR5 exon 1 or regions encoding critical functional domains (N-terminus, ECL2).
- Delivery system: Use lentiviral vectors or electroporation to introduce CRISPR/Cas9 components into primary CD4+ T cells or hematopoietic stem/progenitor cells (HSPCs).
- Editing validation:
- Tracking of Indels by Decomposition (TIDE): PCR amplify target region (Days 3-5 post-editing); sequence and analyze decomposition patterns.
- Flow cytometry: Stain cells with anti-CCR5-APC (clone 2D7) at Day 7 post-editing; quantify CCR5-negative population.
- Functional challenge: Infect edited cells with R5-tropic HIV-1 at Day 10; measure p24 production over 14 days.
- Off-target assessment: Perform whole-genome sequencing or targeted sequencing of predicted off-target sites [3].
Diagram Title: CCR5 Gene Editing Workflow
Therapeutic Targeting of CCR5
Small Molecule Antagonists
Maraviroc (Pfizer), the only FDA-approved CCR5 antagonist, binds within a transmembrane pocket formed by helices I, II, III, VI, and VII, stabilizing CCR5 in an inactive conformation that cannot be engaged by gp120 [4]. Clinical trials demonstrated that maraviroc, when combined with standard antiretroviral therapy, effectively reduces viral load in treatment-experienced patients with R5-tropic virus [2]. Other developed antagonists (vicriviroc, aplaviroc) showed efficacy but faced developmental challenges including toxicity concerns [4].
Gene Editing Approaches
CCR5 gene editing represents a promising therapeutic strategy for achieving HIV-1 functional cure:
- Zinc Finger Nucleases (ZFNs): SB-728-T clinical trials demonstrated that ZFN-modified autologous CD4+ T cells could be safely administered and confer virological benefits, with sustained engraftment of CCR5-disrupted cells [3].
- CRISPR/Cas9: Early-phase clinical trials (NCT03164135) have assessed CRISPR/Cas9-mediated CCR5 editing in hematopoietic stem cells for patients with HIV and acute lymphoblastic leukemia, demonstrating feasibility and safety [3].
- Multiplexed editing strategies: Simultaneous targeting of CCR5, CXCR4, and HIV LTR regions establishes comprehensive viral blockade to counteract potential tropism switching and latent reactivation [3].
Antibody-Based Interventions
PRO 140 (leronlimab), a humanized monoclonal antibody targeting CCR5 ECL2, demonstrates potent antiviral activity by allosterically inhibiting gp120 binding without triggering pro-inflammatory signaling [4]. Phase III clinical trials showed comparable efficacy to daily antiretroviral regimens with weekly subcutaneous administration [4].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for CCR5 Investigation
| Reagent Category | Specific Examples | Research Application | Mechanism/Utility |
|---|---|---|---|
| CCR5 antagonists | Maraviroc, Vicriviroc | Coreceptor inhibition studies | Allosterically stabilizes inactive CCR5 conformation [4] |
| Anti-CCR5 antibodies | 2D7 (neutralizing), 3A9 (sulfation-specific) | Flow cytometry, inhibition assays | Detect surface expression; block specific functional domains [1] [2] |
| Natural ligands | RANTES, MIP-1α, MIP-1β | Competition experiments; signaling studies | Competitive inhibition of HIV-1 entry; study receptor activation [5] [4] |
| Gene editing tools | CRISPR/Cas9 (CCR5 sgRNA), ZFNs | CCR5 disruption studies | Permanent knockout of CCR5 function [3] |
| Cell lines | PM1, U87.CD4.CCR5, TZM-bl | Viral entry and inhibition assays | Stably express CD4 and CCR5; optimized for HIV-1 infection [2] |
Future Directions and Challenges
The continued development of CCR5-targeted therapies faces several important challenges and opportunities:
- Addressing viral tropism switching: CCR5 disruption may select for CXCR4-using (X4-tropic) variants, necessitating combinatorial approaches targeting both coreceptors [3].
- Enhancing safety profiles: Improving specificity of gene editing tools to minimize off-target effects remains a priority for clinical translation [3].
- Economic feasibility: Reducing costs of advanced therapies like gene editing is crucial for global accessibility [3].
- Synergistic immunotherapies: Combining CCR5 editing with broadly neutralizing antibodies, checkpoint inhibitors, or CAR-T cells may more effectively target latent reservoirs [3].
The integration of structural biology insights with innovative therapeutic modalities positions CCR5 at the forefront of HIV-1 cure research, offering promising avenues for overcoming current limitations in antiretroviral therapy and moving toward functional cure strategies.
The fusion of the human immunodeficiency virus type 1 (HIV-1) with host cells represents a critical initial step in viral infection, mediated by a highly coordinated interaction between the viral envelope glycoprotein (Env) and specific host cell receptors. This whitepaper delineates the molecular mechanism of HIV-1 fusion, focusing on the essential cooperative roles of the primary receptor CD4 and the chemokine coreceptor CCR5. The process involves sequential, conformational changes in Env triggered by receptor binding, culminating in the merger of viral and cellular membranes. Within this context, the natural CCR5Δ32 mutation, which confers resistance to HIV-1 infection, is examined alongside therapeutic strategies targeting this mechanism. The discussion is supported by recent structural biology insights, quantitative experimental data, and detailed methodologies relevant to researchers and drug development professionals.
HIV-1 entry into host cells is initiated by the viral Env spike, a trimer of gp120-gp41 heterodimers, binding to the CD4 receptor on the target cell surface [9]. This engagement triggers a series of conformational changes that allow Env to subsequently bind to a coreceptor, predominantly the C-C chemokine receptor type 5 (CCR5), which is utilized by the virus strains (R5-tropic) responsible for the majority of transmissions and prevalent during the early and chronic phases of infection [10] [11]. The cooperative interaction between CD4 and CCR5 is pivotal for activating the fusogenic potential of gp41, leading to the fusion of the viral and cellular membranes and the delivery of the viral core into the cytoplasm. The critical nature of CCR5 is highlighted by the natural CCR5Δ32 mutation, a 32-base-pair deletion that results in a truncated protein not expressed on the cell surface. Individuals homozygous for this mutation are highly resistant to infection by R5-tropic HIV-1 [12] [11]. This review synthesizes the current mechanistic understanding of the CD4-CCR5 cooperative interaction, its role in fusion, and the implications for therapeutic intervention.
Molecular Mechanism of CD4 and CCR5 Cooperation
The HIV-1 fusion process is a cascade of molecular events driven by sequential receptor binding and dramatic structural rearrangements of the Env glycoprotein.
Initial CD4 Binding and Conformational Priming
The attachment begins with the gp120 subunit of Env binding to the D1 domain of the CD4 receptor [9]. CD4 binding destabilizes the metastable, pre-fusion conformation of the Env trimer. Structural studies using cryo-electron microscopy (cryo-EM) reveal that this interaction causes an approximately 40 Å displacement of the V1V2 loops from the apex of the Env trimer, exposing and forming the bridging sheet domain, which is critical for the subsequent coreceptor engagement [13] [14]. This step acts as a conformational "primer," positioning Env for coreceptor binding but not yet triggering membrane fusion.
Coreceptor Engagement and Fusion Activation
The exposed bridging sheet and the tip of the V3 loop of gp120 then engage with the CCR5 coreceptor. The first high-resolution structure of the gp120-CD4-CCR5 complex, solved by cryo-EM, shows that the V3 loop inserts into the extracellular crevice of CCR5, while the CCR5 N-terminus interacts with the gp120 bridging sheet [13]. Unlike CD4 binding, which induces major conformational shifts, CCR5 binding appears to primarily stabilize the CD4-bound state of gp120 and pull the entire complex closer to the target cell membrane [13]. This cooperative binding is highly specific; the conformational changes induced in Env by CD4 are tropism-dependent, meaning that T-cell-tropic Env undergoes significant changes primarily on CD4+ T cells, while macrophage-tropic Env does so more efficiently on CD4+ macrophages [15].
Membrane Fusion Execution
The cooperative engagement of CD4 and CCR5 unleashes the fusogenic potential of the gp41 subunit. Prior to receptor binding, gp41 is held in a metastable state. The dual receptor binding releases constraints on gp41, allowing its hydrophobic fusion peptide (FP) to insert into the target cell membrane. Subsequently, gp41 refolds into a highly stable six-helix bundle (6-HB), where the heptad repeat regions 1 and 2 (HR1 and HR2) form a trimer-of-hairpins. This refolding process mechanically pulls the viral and cellular membranes into close apposition, overcoming the kinetic barriers to membrane merger and pore formation [9].
Table 1: Key Domains and Structural Elements in HIV-1 Fusion
| Component | Key Element | Function in Fusion Process |
|---|---|---|
| gp120 | V1V2 Loops | Shield the coreceptor binding site in pre-fusion state; displaced upon CD4 binding. |
| V3 Loop | Directly inserts into the CCR5 binding pocket; determines coreceptor tropism. | |
| Bridging Sheet | Formed after CD4 binding; critical for stable engagement with CCR5. | |
| gp41 | Fusion Peptide (FP) | Inserts into target cell membrane upon activation. |
| Heptad Repeat 1 (HR1) | Forms the central trimeric coil in the postfusion six-helix bundle. | |
| Heptad Repeat 2 (HR2) | Wraps around the HR1 trimer in the postfusion state. | |
| CCR5 | N-terminal Domain | Interacts with the gp120 bridging sheet. |
| Extracellular Loops | Form a binding pocket for the V3 loop. |
Recent cryo-electron tomography (cryo-ET) studies of Env interacting with membrane-embedded CD4 have revealed that this process is asymmetric and progressive. Env trimers can initially engage one or two CD4 molecules, forming detectable intermediates before the full engagement of three CD4 molecules and the complete opening of the trimer [14]. This clustering of Env-CD4-CCR5 complexes at the virus-cell interface brings the membranes closer together and is dependent on viral capsid maturation, which allows Env the lateral mobility required for clustering [14].
The Role of CCR5 and the CCR5Δ32 Mutation
CCR5 as a Paradigm for HIV-1 Host Dependency Factors
CCR5 is a G-protein coupled receptor (GPCR) expressed on various immune cells, including memory/effector T lymphocytes, macrophages, and dendritic cells [11]. Its natural ligands are chemokines such as CCL3 (MIP-1α), CCL4 (MIP-1β), and CCL5 (RANTES), which are involved in immune cell recruitment [10] [11]. HIV-1 exploits this receptor as a primary entry portal. The level of CCR5 expression correlates with cellular susceptibility to infection, and the receptor's conformation is dynamically regulated by post-translational modifications like sulfation of tyrosine residues in its N-terminus, which enhances gp120 binding [1].
The Protective CCR5Δ32 Mutation
The CCR5Δ32 allele contains a 32-base-pair deletion that causes a frameshift and premature stop codon, resulting in a truncated protein that is not expressed on the cell surface [11]. This loss-of-function mutation has profound implications for HIV-1 resistance. A meta-analysis of 24 case-control studies demonstrated that individuals who are homozygous (delta32/delta32) have a significantly reduced susceptibility to HIV-1 infection (OR=0.25, 95%CI=0.09-0.68) [12]. Conversely, heterozygous carriers may have a slightly increased susceptibility (OR=1.16, 95%CI=1.02-1.32), though they often experience a slower disease progression, suggesting a gene dosage effect where reduced CCR5 expression is beneficial but not fully protective [12]. The mutation is most prevalent in Northern European populations, with a frequency of up to 16% in Finland and Russia, and is virtually absent in African, Asian, and Native American populations [11].
Table 2: Impact of CCR5Δ32 Genotype on HIV-1 Susceptibility
| Genotype | Receptor Expression | Effect on HIV-1 Susceptibility | Odds Ratio (OR) vs. Wild-Type |
|---|---|---|---|
| Wild-Type Homozygous | Normal | Normal susceptibility | Reference (OR=1) |
| Heterozygous | Reduced | Slightly increased susceptibility / Slower disease progression | OR=1.16 (95%CI: 1.02-1.32) |
| Δ32 Homozygous | Not detected | Strongly reduced susceptibility | OR=0.25 (95%CI: 0.09-0.68) |
The "Berlin" and "London" patients, who were cured of HIV after receiving hematopoietic stem cell transplants from CCR5Δ32 homozygous donors, provide compelling clinical validation of CCR5 as a therapeutic target [3].
Advanced Signaling and Enhancement Mechanisms
Beyond acting as a passive docking site, CCR5 actively participates in signaling that enhances the fusion process. Engagement of gp120 with CCR5 triggers intracellular Ca2+ signaling that activates the lipid scramblase TMEM16F [16]. This leads to the externalization of the membrane phospholipid phosphatidylserine (PS) to the outer leaflet of the target cell membrane. This exposed PS acts as a critical cofactor for HIV-1 entry, strongly promoting Env-mediated membrane fusion. Blocking PS with binding proteins or inhibiting TMEM16F suppresses fusion, while adding exogenous PS enhances it, an effect particularly pronounced in cells with low coreceptor density [16]. This represents a bi-directional signaling pathway where outside-in signaling through CCR5 triggers inside-out PS signaling to facilitate infection.
Diagram 1: HIV-1 Fusion and Signaling Pathway
Experimental Methods and Research Tools
Understanding the HIV-1 fusion mechanism relies on a suite of sophisticated biochemical, structural, and cell biological techniques.
Key Experimental Protocols
1. Cell-Cell Fusion Assay: This is a common method to study Env-mediated membrane fusion. Effector cells expressing HIV-1 Env (e.g., HeLa-Env cells) are co-cultured with target cells expressing CD4 and CCR5 (e.g., TZM-bl cells). Fusion is quantified by the cytoplasmic mixing of fluorescent markers (e.g., eGFP and mCherry) or reporter gene activation. This assay allows for the testing of fusion inhibitors (e.g., TAK-779, C52L peptide) and enhancers (e.g., exogenous phosphatidylserine) [16].
2. Cryo-Electron Tomography (Cryo-ET): This technique allows for the high-resolution visualization of Env-receptor interactions in a native membrane context. Researchers incubate HIV-1 virions with CD4-presenting virus-like particles (VLPs) and plunge-freeze the mixture. Cryo-ET reconstructions reveal the spatial organization of Env-CD4 complexes, showing their clustering and the progressive binding of one, two, and three CD4 molecules per trimer, providing direct evidence for asymmetric intermediate states [14].
3. PS Externalization Measurement: The role of phosphatidylserine is studied using fluorescently labeled probes like the C2 domain of lactadherin (LactC2), which specifically binds externalized PS. Flow cytometry or fluorescence microscopy can then quantify and visualize PS exposure on target cells triggered by HIV-1 Env or gp120 binding. The specific involvement of TMEM16F is validated using pharmacological inhibitors (e.g., CaCCinh-A01) or RNAi-mediated knockdown [16].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Studying HIV-1 Fusion and CCR5 Interaction
| Reagent / Tool | Type | Function and Application |
|---|---|---|
| SOSIP Trimers | Stabilized Env Protein | Soluble, native-like Env trimers used for structural studies (cryo-EM, X-ray) and immunization. |
| TZM-bl Cells | Reporter Cell Line | HeLa-derived cells expressing high levels of CD4 and CCR5; contain reporter genes for quantifying infection/fusion. |
| TAK-779 | Small Molecule Antagonist | Specifically blocks gp120 binding to CCR5; used to validate CCR5-dependent fusion. |
| AMD-3100 (Plerixafor) | Small Molecule Antagonist | Specifically blocks gp120 binding to CXCR4; used as a control and to study X4-tropic viruses. |
| Lactadherin (LactC2) | PS-binding Protein | Binds to externalized phosphatidylserine; used to detect and block PS-enhanced fusion. |
| CaCCinh-A01 | Small Molecule Inhibitor | Inhibits TMEM16F scramblase activity; used to probe the role of PS externalization. |
| C52L Peptide | Fusion Inhibitor | Peptide derived from gp41 HR2 that inhibits the formation of the six-helix bundle. |
Diagram 2: Fusion and Structural Analysis Workflows
Therapeutic Implications and Future Directions
The detailed understanding of the CD4-CCR5 fusion mechanism has directly informed the development of entry inhibitors.
1. CCR5 Antagonists: Drugs like maraviroc bind directly to the CCR5 transmembrane pocket and sterically hinder the interaction with the gp120 V3 loop, as confirmed by structural studies [13]. This class of drugs is a cornerstone for treating patients with R5-tropic virus.
2. Fusion Inhibitors: Enfuvirtide (T-20) is a peptide derived from HR2 of gp41 that disrupts the formation of the six-helix bundle, preventing the final step of membrane fusion [9].
3. Gene Editing Strategies: Inspired by the natural resistance of CCR5Δ32 homozygotes, CRISPR/Cas9 and other nuclease technologies are being explored to disrupt the CCR5 gene in patients' hematopoietic stem cells or T cells, aiming for a functional cure [3]. Current research focuses on multiplexed editing strategies that simultaneously target CCR5, CXCR4, and the viral LTR to prevent coreceptor switching and viral reactivation from latency [3].
4. Antibody-Based Therapies: Broadly neutralizing antibodies (bNAbs) that target the Env trimer can block CD4 or coreceptor binding sites. Furthermore, the natural occurrence of anti-CCR5 antibodies in some exposed uninfected individuals has prompted research into CCR5-targeted vaccinogen designs [10].
The cooperative interaction between CD4 and CCR5 is a finely orchestrated process that is fundamental to HIV-1 entry. The mechanism proceeds through defined stages: CD4 binding and priming, asymmetric coreceptor engagement, intracellular signaling, and final execution of fusion via gp41 refolding. The protective effect of the CCR5Δ32 mutation and the clinical efficacy of CCR5 antagonists validate this pathway as a critical therapeutic target. Future efforts combining structural biology, gene editing, and immunology hold the promise of developing more effective interventions, moving closer to the ultimate goals of prevention and cure.
The C-C chemokine receptor 5 (CCR5) is a seven-transmembrane, G protein-coupled receptor (GPCR) primarily expressed on immune cells including lymphocytes, macrophages, and dendritic cells [17]. Its normal physiological role involves orchestrating immune cell migration (chemotaxis) to sites of inflammation along chemokine gradients of its natural ligands CCL3, CCL4, and CCL5 [17]. However, CCR5 gained significant scientific notoriety for its role as the primary co-receptor for human immunodeficiency virus type 1 (HIV-1) during initial transmission and early infection stages [18] [17]. HIV-1 entry into CD4+ T-cells requires sequential binding of the viral envelope glycoprotein gp120 to the CD4 receptor followed by interaction with either CCR5 or CXCR4 co-receptors [19]. Virus strains are classified as R5-tropic (CCR5-using), X4-tropic (CXCR4-using), or dual-tropic based on this coreceptor preference [19]. The CCR5-Δ32 genetic variant, a 32-base-pair deletion in the CCR5 gene, produces a truncated, non-functional receptor that fails to localize to the cell surface, thereby conferring strong resistance to HIV-1 infection in homozygous individuals [20] [17]. This natural resistance mechanism has inspired novel therapeutic strategies aimed at curing HIV-1 through CCR5 disruption [21] [18] [22].
Molecular Genetics and Protein Structure of CCR5-Δ32
Genetic Architecture and Mutational Effect
The CCR5-Δ32 variant is characterized by a 32-base-pair deletion (Δ32) within the coding region of the CCR5 gene on chromosome 3p21.31 [20]. This deletion localizes to the region encoding the second extracellular loop of the receptor [17] and introduces a premature stop codon through frameshift, resulting in a truncated protein that is severely impaired in structure and function [20]. The mutant protein does not express on the cell surface due to retention in the endoplasmic reticulum and accelerated degradation [19] [20]. Recent genetic studies reveal that the CCR5Δ32 deletion is part of a specific haplotype (Haplotype A) comprising 86 linked variants in high linkage disequilibrium, with two single nucleotide polymorphisms (rs113341849 and rs113010081) in perfect LD with CCR5Δ32 [20].
Structural Consequences for the CCR5 Receptor
The structural impact of the Δ32 mutation is profound. The normal CCR5 receptor contains seven transmembrane α-helices with extracellular and intracellular domains critical for its function [17]. The deletion occurs just before the third extracellular element (second loop) that contains the essential 2D7 HIV binding site [23]. Consequently, the mutant protein lacks this critical structural domain while retaining the PA12 binding site [23]. This structural ablation prevents the conformational changes required for HIV-1 gp120 binding and subsequent viral fusion, thereby blocking viral entry into target cells [23] [17].
Table 1: Molecular Characteristics of Wild-Type CCR5 vs. CCR5-Δ32 Mutant
| Feature | Wild-Type CCR5 | CCR5-Δ32 Mutant |
|---|---|---|
| Gene Length | Full-length CCR5 gene | 32-bp deletion in coding region |
| Protein Structure | Seven transmembrane domains with intact extracellular loops | Truncated protein missing critical extracellular domains |
| Cellular Localization | Cell surface expression | Retained intracellularly, not expressed on surface |
| HIV Binding Site (2D7) | Present and functional | Absent due to deletion |
| Receptor Function | Functional chemokine receptor and HIV-1 co-receptor | Non-functional for both chemokine signaling and HIV-1 entry |
Population Genetics and Evolutionary History
Global Distribution and Ethnic Variation
The CCR5-Δ32 allele demonstrates a striking geographical distribution pattern, predominantly found in European Caucasian populations with an average frequency of approximately 10% and homozygosity frequency of about 1% [20] [17]. The frequency displays marked North-South and East-West clines across Europe, with highest frequencies observed in Nordic countries (16% in Finland and Russia, 15% in Iceland, 14% in Sweden) and lowest frequencies in Southern Europe (4% in Sardinia, 0.9% in Corsica) [17]. The allele is virtually absent in native African, Asian, and Native American populations [20] [17], suggesting a relatively recent origin after the divergence of European populations.
Evolutionary Origins and Selective Pressure
Multiple lines of evidence indicate the CCR5-Δ32 mutation arose from a single mutational event [20] [24]. Genetic linkage analyses demonstrate the mutation occurs on a homogeneous genetic background, with over 95% of CCR5-Δ32 chromosomes carrying identical flanking microsatellite markers [20]. Age estimates for the mutation vary between 700-2,500 years, significantly younger than the time required to reach current frequencies through genetic drift alone [20] [17]. This discrepancy indicates the allele underwent intense positive selection. While bubonic plague (Yersinia pestis) was initially proposed as the selective agent, recent evidence stronger supports smallpox (Variola major) as the driving selective pressure due to its longer historical presence, higher childhood mortality, and viral mechanism potentially exploiting CCR5 similar to HIV-1 [20]. The Viking dispersals in the 8th-10th centuries may have facilitated the spread of this mutation throughout Europe [20] [17].
Table 2: Global Distribution of CCR5-Δ32 Allele Frequency
| Population | Region | Δ32 Allele Frequency (%) | Homozygous Frequency (%) |
|---|---|---|---|
| Northern European | Finland, Russia | 16 | ~2.6 |
| Scandinavian | Iceland, Sweden, Denmark | 13-15 | ~1.7-2.3 |
| Western European | Northern France, Germany | 10-14 | ~1-2 |
| Southern European | Spain, Italy, Portugal | 5-7 | ~0.3-0.5 |
| Mediterranean | Sardinia, Corsica | 0.9-4 | ~0.01-0.16 |
| African | Sub-Saharan Africa | Virtually absent | 0 |
| Asian | East Asia | Virtually absent | 0 |
| Native American | Americas | Virtually absent | 0 |
Protective Effects Against HIV-1 Infection
Resistance Mechanisms and Genotype-Phenotype Correlation
The CCR5-Δ32 mutation confers resistance to HIV-1 through multiple complementary mechanisms. In homozygous individuals (Δ32/Δ32), the complete absence of functional CCR5 receptors on the cell surface prevents R5-tropic HIV-1 strains from entering target CD4+ T-cells [20] [17]. Heterozygous individuals (+/Δ32) exhibit approximately 50% reduction in surface CCR5 expression due to dimerization between mutant and wild-type receptors that interferes with normal receptor trafficking to the cell membrane [20]. Additionally, the CCR5-Δ32 protein can form heterodimers with wild-type CCR5 and CXCR4, leading to intracellular retention and reduced surface expression of both coreceptors [19]. Recent research also indicates that CD4+ T lymphocytes isolated from CCR5-Δ32 homozygous individuals show lower levels of CXCR4 expression that correlate with reduced X4 Env-mediated fusion, providing additional protection against CXCR4-tropic viruses [19].
Clinical Evidence and Meta-Analysis Findings
A comprehensive meta-analysis of 24 case-control studies involving 4,786 HIV-1 patients and 6,283 controls quantified the protective effects of CCR5-Δ32 [12]. Compared to wild-type homozygous individuals, CCR5-Δ32 heterozygotes had slightly increased susceptibility to HIV-1 acquisition (OR=1.16, 95% CI=1.02-1.32), potentially due to risk-behavior compensation or other confounding factors [12]. However, once infected, heterozygous individuals demonstrated delayed disease progression and reduced viral loads [20]. Most significantly, Δ32 homozygotes showed substantially reduced susceptibility to HIV-1 infection (OR=0.25, 95% CI=0.09-0.68) compared to healthy controls [12]. When using HIV-exposed but uninfected individuals as controls, Δ32 allele carriers showed significantly reduced HIV-1 susceptibility (OR=0.71, 95% CI=0.54-0.94) [12]. Rare cases of HIV-1 infection in CCR5-Δ32 homozygous individuals have been reported, typically involving X4-tropic or dual-tropic viruses [19]. Investigations of these rare cases revealed that some infected homozygotes exhibited absent or reduced CCR5-Δ32 protein expression, suggesting additional factors beyond genotype alone determine HIV-1 resistance [19].
Experimental Methodologies and Research Applications
Key Experimental Protocols for CCR5-Δ32 Research
Genotyping and Expression Analysis
Flow Cytometry for Surface Expression: Peripheral blood mononuclear cells (PBMCs) are washed twice in FACS buffer and resuspended at 10^7/ml concentration. Cells are incubated with 1:200 dilution of fluorescently-labeled monoclonal antibodies against CD4, CCR5 (clone 556042), and CXCR4 (clone 555976) at 4°C for 30 minutes [19]. Sequential staining followed by washing steps enables multi-parameter analysis of coreceptor expression. Cells are analyzed using a FACScan cytometer, with CCR5 expression quantified on CD4+ T-cell populations [19].
HIV-1 Env-Mediated Cell Fusion Assay: This assay evaluates functional coreceptor activity using two cell populations [19]. Target PBMCs (expressing endogenous CD4) are infected with vCB-21R (encoding LacZ under T7 promoter). HeLa cells coinfected with vTF7-3 and HIV-1 Env serve as effector cells. After mixing effector and target populations and incubating at 37°C for 2.5 hours, fusion specificity is measured by β-galactosidase production in a colorimetric lysate assay [19].
Western Blotting for Protein Detection: Cell lysates from PBMCs or cell lines are prepared and fractionated on 12.5% SDS-polyacrylamide gel electrophoresis [19]. Immunoblotting verifies expression of endogenous or adenovirus-encoded CCR5-Δ32 protein, with specific antibodies detecting the truncated form compared to wild-type CCR5.
CCR5 Gene Editing using CRISPR/Cas9
Recent advances enable precise CCR5 ablation using CRISPR/Cas9 technology [22]. The optimized protocol involves:
Guide RNA Selection: In silico prediction identifies gRNAs targeting exon 3 of CCR5. After excluding gRNAs with potential off-target effects, optimal guides (e.g., TB48, TB50) are selected based on editing efficiency (>90% in HSPCs) and minimal off-target activity [22].
Electroporation of HSPCs: Mobilized human CD34+ hematopoietic stem progenitor cells are electroporated with Cas9 protein complexed with gRNAs as ribonucleoproteins (RNPs) [22].
Transplantation and Engraftment: Edited HSPCs are transplanted into immunodeficient mice, demonstrating normal hematopoiesis and production of CCR5 null T-cells that confer refractoriness to HIV infection [22].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for CCR5-Δ32 Investigation
| Reagent/Category | Specific Examples | Research Application | Technical Notes |
|---|---|---|---|
| Cell Lines | PBMCs from genotyped donors, HEK293T, HeLa cells | Functional fusion assays, viral entry studies | Primary cells best reflect physiological expression |
| Antibodies | Anti-CCR5 (clone 556042), Anti-CXCR4 (clone 555976), Anti-CD4 (clone 555346) | Flow cytometry, Western blotting | Conjugated with PE, FITC, APC for multi-color FACS |
| Gene Editing Tools | CRISPR/Cas9 (gRNAs TB48, TB50), ZFNs, TALENs | CCR5 ablation studies, therapeutic development | CRISPR/Cas9 RNP delivery shows >90% editing in HSPCs |
| Viral Vectors | Adenovirus 5/Δ32, HIV-1 JRCSF (R5-tropic) | Protein expression, viral challenge assays | Ad5-encoded CCR5-Δ32 used for functional rescue studies |
| Assay Systems | HIV-1 Env-mediated fusion assay, p24 ELISA | Viral entry and replication quantification | β-galactosidase reporter system for fusion efficiency |
Therapeutic Applications and Clinical Translation
CCR5-Targeted Gene Therapy Approaches
The successful cure of HIV-1 in the "Berlin," "London," and "Düsseldorf" patients through transplantation with CCR5-Δ32/Δ32 hematopoietic stem cells demonstrated the therapeutic potential of CCR5 disruption [21] [22]. This proof-of-concept has inspired the development of autologous stem cell therapies using gene editing to recreate the CCR5-Δ32 phenotype in patients' own cells [22]. Multiple gene editing platforms have been employed:
CRISPR/Cas9 System: The most widely used approach employs Cas9 nuclease complexed with guide RNAs (e.g., TB48, TB50) to introduce double-strand breaks in the CCR5 locus [22]. Recent optimized protocols achieve >90% editing efficiency in human hematopoietic stem progenitor cells (HSPCs) while maintaining normal pluripotency and engraftment capacity [22]. Titration studies demonstrate that >90% CCR5 editing is required for consistent protective benefit, with lower editing frequencies (54-26%) providing diminishing protection [22].
Zinc Finger Nucleases (ZFNs): Among the earliest technologies applied clinically, ZFNs use custom-designed zinc finger proteins fused to FokI nuclease domains [21] [3]. The SB-728-T clinical trial demonstrated that autologous T-cells edited by ZFNs and reinfused into patients yielded acceptable safety profiles and virological benefits [3].
TALENs and Base Editors: Transcription activator-like effector nucleases (TALENs) offer improved specificity over ZFNs, while base editors enable precise nucleotide conversions without double-strand breaks, minimizing risks of indels and chromosomal translocations [21] [3].
Multi-Target Strategies and Combinatorial Approaches
Single-target CCR5 editing faces limitations due to potential viral tropism switching to CXCR4 usage [21]. To address this, multiplexed gene editing strategies simultaneously target CCR5, CXCR4, and HIV long terminal repeat (LTR) regions [21] [3]. This comprehensive approach establishes multiple barriers to viral replication by preventing entry via both major coreceptors and suppressing viral reactivation from latent reservoirs [21]. Additionally, combinatorial approaches integrating gene editing with immunotherapy show promise, such as engineering HIV-specific CAR-T cells with edited CCR5 genes or checkpoint inhibitor expression to enhance both resistance to infection and viral clearance capacity [21] [3].
The CCR5-Δ32 mutation represents a remarkable example of human evolution responding to historical pathogenic pressures, now providing profound insights for modern therapeutic development. The molecular characterization of this 32-bp deletion and its profound effect on HIV-1 resistance has illuminated fundamental aspects of viral entry mechanisms while inspiring innovative gene editing strategies. Current research focuses on optimizing editing efficiency, ensuring long-term safety, and developing accessible delivery platforms to translate this natural protective mechanism into broadly applicable HIV-1 therapies. The continued investigation of CCR5-Δ32 biology not only advances HIV-1 cure efforts but also provides valuable insights into chemokine receptor function, host-pathogen interactions, and the development of precision genetic medicines for infectious diseases.
The CCR5-Δ32 allele, a 32-base pair deletion in the CC chemokine receptor type 5 (CCR5) gene, represents a paradigm in human population genetics and evolutionary biology. This allele, which confers resistance to HIV-1 infection in homozygous individuals, exhibits a unique and restricted geographic distribution that has fueled extensive scientific inquiry into its origins. This review synthesizes current understanding of the allele's global distribution patterns, evolutionary history, and the methodological frameworks employed in its study. We examine the evidence supporting various selective pressures that may have driven its frequency increase in European populations, from historical pandemics to other pathogenic challenges. Furthermore, we detail the experimental protocols essential for genetic screening and functional analysis of CCR5-Δ32, providing researchers with a technical toolkit for continued investigation. The discussion is situated within the broader context of CCR5's role as an HIV co-receptor and the implications of CCR5-Δ32 research for therapeutic development.
CCR5 is a G-protein-coupled receptor (GPCR) expressed on the surface of various immune cells, including T lymphocytes, macrophages, and dendritic cells [25] [11]. Its natural ligands include chemokines such as CCL3 (MIP-1α), CCL4 (MIP-1β), and CCL5 (RANTES), which stimulate cell migration and mediate inflammatory responses [25] [11]. Beyond its immunological functions, CCR5 serves as the principal co-receptor for human immunodeficiency virus type 1 (HIV-1) entry into CD4+ T cells [11]. The virus exploits the interaction between its envelope glycoprotein gp120, the CD4 receptor, and CCR5 to gain cellular entry, making this receptor critical for the initial stages of HIV infection [11].
The CCR5-Δ32 genetic variant is a 32-base pair deletion in the CCR5 gene's coding region that causes a frameshift mutation, resulting in a truncated protein that is not expressed on the cell surface [25] [11]. Individuals homozygous for this mutation lack functional CCR5 receptors on their cell surfaces and exhibit substantial resistance to infection with CCR5-tropic (R5) HIV-1 strains [25] [12]. Heterozygous individuals show reduced CCR5 expression and may experience slower disease progression if infected [25]. This protective effect has positioned CCR5-Δ32 as a focal point in HIV research, inspiring therapeutic strategies ranging from small-molecule antagonists like Maraviroc to innovative gene-editing approaches [25] [26] [11].
Despite its significance in HIV pathogenesis, the CCR5-Δ32 allele exists at appreciable frequencies primarily in European populations, with much lower frequencies or complete absence in other geographic regions [11] [24]. This unusual distribution pattern suggests a complex evolutionary history, potentially involving selective pressures that predate the HIV pandemic. This review examines the population genetics of the CCR5-Δ32 allele, exploring its global distribution, evolutionary origins, and the experimental methodologies that underpin its study.
Molecular Genetics and Functional Impact of CCR5-Δ32
Molecular Structure and Consequences of the Deletion
The CCR5 gene is located on chromosome 3 (3p21.31) and encodes a protein of 352 amino acids [25]. The protein structure comprises seven transmembrane α-helices, three extracellular loops, three intracellular loops, an amino-terminal domain, and a carboxyl-terminal domain [11]. The CCR5-Δ32 deletion occurs in a region encoding the second extracellular loop of the receptor, which is critical for chemokine binding and HIV coreceptor function [11].
The deletion causes a frameshift mutation that generates a premature stop codon, resulting in a severely truncated protein of only 215 amino acids instead of the full-length 352-amino-acid protein [25] [27]. This truncated protein is retained intracellularly and degraded, preventing its expression on the cell surface [25]. Consequently, homozygous individuals effectively lack CCR5 on their cell surfaces, creating a barrier to cellular entry for R5-tropic HIV-1 strains [25].
Functional Impacts on HIV-1 Infection
The absence of functional CCR5 receptors in homozygous individuals provides nearly complete protection against infection with CCR5-tropic HIV-1 strains, which are responsible for the majority of transmissions [25] [12]. Heterozygous individuals display reduced CCR5 expression on their cell surfaces, which does not prevent infection but can delay disease progression to AIDS [12] [27]. This gene dosage effect underscores the critical role of CCR5 expression levels in HIV susceptibility and pathogenesis.
The diagram below illustrates the molecular consequences of the CCR5-Δ32 mutation and its impact on HIV infection:
Beyond HIV infection, the CCR5-Δ32 variant modifies CCR5-mediated inflammatory responses in various conditions [25] [28]. Research indicates that this polymorphism may influence susceptibility to other infectious diseases, including West Nile virus, influenza virus, and flaviviruses, though these effects vary significantly between pathogens [25] [28]. For instance, the Δ32 allele increases the risk of symptomatic West Nile virus infection while potentially conferring protective effects against other pathogens [25].
Global Distribution Patterns
The CCR5-Δ32 allele demonstrates a distinctive and uneven global distribution, with highest frequencies observed in European populations, particularly those of Northern European descent [25] [11]. The table below summarizes the allele's frequency across different global populations:
Table 1: Global Distribution of CCR5-Δ32 Allele Frequencies
| Region/Country | Population Description | CCR5-Δ32 Frequency | Source |
|---|---|---|---|
| Northern Europe | |||
| Norway | General population | 16% | [25] |
| Finland & Russia | General population | 16% | [11] |
| Iceland | General population | 15% | [11] |
| Sweden | General population | 14% | [11] |
| Denmark | General population | 13% | [11] |
| Western Europe | |||
| Germany | General population | 11% | [25] |
| Northern France | General population | 14% | [11] |
| Luxembourg | General population | ~10% (estimated) | [12] |
| Southern Europe | |||
| Spain | General population | 7% | [11] |
| Italy | General population | 5.6% | [11] |
| Portugal | General population | 5.2% | [11] |
| Sardinia | General population | 4% | [11] |
| Corsica | General population | 0.9% | [11] |
| Other Regions | |||
| South Africa | European-derived | 13% | [25] [11] |
| Chile | European-admixed | 12% | [25] [11] |
| Brazil | General population | 4-5% | [25] |
| Brazil | Southern region | 9% | [25] |
| United States | African American | 2% | [11] |
| Angola | General population | 0% | [29] |
| Peru | General population | 2.7% (heterozygous) | [30] |
| India | National Capital Regions | 2.8% | [27] |
This distribution pattern reveals a pronounced north-south cline within Europe, with frequencies highest in Scandinavian and Nordic populations and gradually decreasing toward Mediterranean regions [11]. The allele is virtually absent in indigenous populations of Sub-Saharan Africa, East Asia, and the Americas, indicating it arose after the divergence of these populations [29] [11] [24].
Population admixture resulting from migration has introduced the Δ32 allele to various regions worldwide. In South Africa, Chilean, and Brazilian populations, the allele frequency correlates with the degree of European ancestry [25]. Similarly, the presence of CCR5-Δ32 in African Americans (approximately 2%) reflects gene flow from European populations [11].
Recent studies continue to refine our understanding of the allele's distribution. A 2025 study from Peru found a CCR5/CCR5-Δ32 heterozygous prevalence of 2.7% with no homozygous individuals detected [30]. The population was in Hardy-Weinberg equilibrium for the CCR5 locus, and no statistical difference in frequency was observed between HIV-exposed seronegative individuals and HIV-seropositive individuals [30]. Similarly, a 2025 study from Angola reported a complete absence of the CCR5-Δ32 allele in their cohort of 272 individuals [29], confirming the allele's rarity in African populations.
Evolutionary Origins and Selective Pressures
Population Genetics and Age Estimates
The CCR5-Δ32 allele is evolutionarily young, with estimates suggesting it arose between 700 and 2000 years ago [24]. Its current high frequencies in European populations, despite this recent origin, indicate strong positive selection acting upon this mutation [24]. The spatial distribution of the allele, with highest frequency in Northern Europe, suggests a potential origin in this region, possibly followed by dispersal by Viking populations during the 8th-10th centuries [11].
The discordance between the allele's young age and its current high frequency presents an evolutionary puzzle. Mathematical models indicate that neutral drift alone is insufficient to explain its rapid increase in frequency, strongly implying the action of historical selective pressure [24].
Proposed Selective Agents
Several hypotheses have been proposed to explain the selective advantage conferred by CCR5-Δ32 in European populations:
1. Historical Plagues: The bubonic plague (Black Death) in the 14th century was initially proposed as a potential selective event, with the hypothesis that CCR5-Δ32 conferred resistance to Yersinia pestis infection. However, subsequent research has yielded conflicting evidence, and this hypothesis remains controversial [24].
2. Smallpox: Variola virus (smallpox) has been suggested as a stronger candidate selective pressure, given its long history in European populations, high mortality rates, and significant impact on population demographics. Some research indicates that the CCR5 receptor may play a role in the immune response to poxviruses, potentially giving Δ32 carriers a survival advantage [24].
3. Other Pathogens: The "pathogen package" hypothesis suggests that multiple consecutive or simultaneous epidemics (e.g., plague, smallpox, typhus, or diphtheria) could have collectively driven the selection for CCR5-Δ32 in European populations [24].
4. Non-Infectious Selective Pressures: Some researchers have proposed that the allele may have provided advantages in specific environmental conditions or nutritional circumstances present in historical Europe, though evidence for these hypotheses remains limited.
The diagram below illustrates the evolutionary timeline and potential selective pressures that have shaped the distribution of the CCR5-Δ32 allele:
The selective advantage hypothesis is further supported by evidence of parallel evolution in nonhuman primate species. Several African monkey species that are natural hosts of simian immunodeficiency virus (SIV) have developed independent mutations affecting CCR5 expression and function, suggesting convergent evolution in response to lentiviral pathogens [11] [24]. These natural strategies include deletion mutations, downregulation of CCR5 expression on CD4+ T cells, and delayed onset of CCR5 expression during ontogenetic development [11].
Methodological Approaches in CCR5-Δ32 Research
Genotyping Techniques
DNA Extraction and Sample Preparation: Genetic studies of CCR5-Δ32 typically begin with DNA extraction from peripheral blood mononuclear cells (PBMCs) or other biological samples. Commercial kits such as the QIAamp DNA Mini Kit (Qiagen) are commonly employed following manufacturer's protocols [29] [30]. For field studies or resource-limited settings, DNA can be extracted from dried blood spots (DBS) on filter paper such as Whatman 903, providing a stable medium for DNA preservation and transportation [29].
PCR Amplification and Genotyping: The primary method for CCR5-Δ32 detection involves conventional polymerase chain reaction (PCR) with primers flanking the 32-bp deletion region. A standard protocol includes:
- Primers: Forward: 5′-ACCAGATCTCTCAAAAAGAAGGTCT-3′ and Reverse: 5′-CATGATGGTGAAGATAAGCCTCCACA-3′ [30]. Alternative primer sequences include Forward: 5′-CTTCATCATCCTCCTGACAATCG-3′ and Reverse: 5′-GACCAGCCCCAAGTTGACTATC-3′ [29].
- Reaction Mix: 5μL of extracted DNA (approximately 1.5-19ng), 12.5μL of GoTaq Green Master Mix (Promega) or NZYTaq II 2x Green Master Mix (NZYTECH), 2.5μL of each primer (0.2μM final concentration), and 2.5μL of ultrapure water for a final volume of 20μL [29].
- Thermocycling Conditions: Initial denaturation at 95°C for 5 minutes; 35 cycles of 95°C for 45 seconds, 58-60°C for 45 seconds, and 72°C for 45 seconds; final extension at 72°C for 10 minutes [29] [30].
- Product Visualization: PCR products are separated by electrophoresis on 2-3% agarose gel and visualized under UV light [29] [30]. Wild-type alleles produce a 225-bp fragment, heterozygous individuals show both 225-bp and 193-bp fragments, while Δ32 homozygotes display only the 193-bp fragment [30].
Alternative Genotyping Methods: More advanced techniques include real-time PCR with specific probes for HLA-B*57:01 genotyping (often studied alongside CCR5-Δ32) [30] and DNA sequencing to confirm novel polymorphisms [27]. Sanger sequencing is typically performed using Big Dye Terminator reagent with analysis on genetic analyzers such as the Applied Biosystems 3500 XL [30].
The workflow for CCR5-Δ32 genotyping is illustrated below:
Functional Assays
Cell Surface Expression Analysis: Flow cytometry is employed to quantify CCR5 expression on PBMCs using fluorescently labeled antibodies against CCR5 (e.g., anti-CCR5 monoclonal antibodies) [25]. Cells are stained and analyzed on flow cytometers to compare expression levels between different genotypes.
Viral Infectivity Assays: In vitro infection assays assess the functional consequences of CCR5-Δ32. Primary CD4+ T cells isolated from individuals of different genotypes are exposed to R5-tropic and X4-tropic HIV-1 strains [26]. Viral replication is monitored by measuring p24 antigen production or viral RNA levels over time [26].
Quantitative Viral Outgrowth Assay (qVOA): This sensitive technique detects latent HIV-1 reservoirs by activating resting CD4+ T cells and measuring virus production [26]. In CCR5-Δ32 research, qVOA has been used to demonstrate the absence of replication-competent virus in patients following CCR5-Δ32 stem cell transplantation [26].
Research Reagent Solutions
Table 2: Essential Research Reagents for CCR5-Δ32 Studies
| Reagent/Category | Specific Examples | Function/Application |
|---|---|---|
| DNA Extraction Kits | QIAamp DNA Mini Kit (Qiagen), NucleoSpin kit (Macherey-Nagel) | High-quality DNA extraction from whole blood, PBMCs, or dried blood spots |
| PCR Master Mixes | GoTaq Green Master Mix (Promega), NZYTaq II 2x Green Master Mix (NZYTECH) | Robust amplification of CCR5 gene fragments |
| Specialized PCR Reagents | Kapa Probe Fast Master Mix (Roche) | Real-time PCR for HLA-B*57:01 genotyping |
| Electrophoresis Materials | Agarose, TBE buffer, DNA size markers, ethidium bromide/SYBR Safe | Separation and visualization of PCR products |
| Sequencing Reagents | Big Dye Terminator reagent, Sequencing primers | Sanger sequencing confirmation of mutations |
| Cell Culture Reagents | RPMI medium, FBS, cytokines (IL-2) | Maintenance and expansion of primary lymphocytes |
| Flow Cytometry Reagents | Anti-CCR5 antibodies, anti-CD4 antibodies, isotype controls | Cell surface receptor quantification |
| Viral Assay Reagents | p24 antigen ELISA kits, viral RNA extraction kits | Measurement of HIV replication |
Clinical Applications and Therapeutic Implications
The understanding of CCR5-Δ32 biology has directly translated into clinical applications. The most successful example is Maraviroc, a CCR5 antagonist approved for HIV treatment that mimics the protective effect of the Δ32 mutation by allosterically blocking the receptor [25] [11]. Additionally, the Berlin Patient (Timothy Ray Brown) and London Patient represent groundbreaking cases of HIV remission following hematopoietic stem-cell transplantation (HSCT) from CCR5-Δ32 homozygous donors [26].
These clinical successes have inspired next-generation therapeutic approaches, including gene editing strategies using technologies like CRISPR-Cas9 to disrupt CCR5 expression in autologous cells [11]. Gene silencing approaches employing RNA interference (RNAi) to downregulate CCR5 expression also show promise [11]. However, given CCR5's role in immune surveillance and response to other pathogens, the potential consequences of long-term CCR5 blockade or elimination require careful consideration [25] [11].
The CCR5-Δ32 allele represents a remarkable example of recent human evolution, with its unique geographic distribution and significant implications for infectious disease resistance. The pronounced north-south gradient in Europe and virtual absence in other indigenous populations continues to fuel research into its evolutionary origins. Methodological advances in genotyping and functional analysis have enabled detailed characterization of this polymorphism and its biological consequences.
The allele's protective effect against HIV infection has not only provided crucial insights into viral pathogenesis but has also inspired novel therapeutic strategies, from small-molecule antagonists to stem-cell transplantation and gene editing approaches. Future research directions include further elucidating the historical selective pressures that shaped the allele's distribution, exploring its implications for other infectious and inflammatory conditions, and developing safer, more precise methods for therapeutic CCR5 modulation.
As genetic research continues to advance, the CCR5-Δ32 allele will undoubtedly remain a focal point for understanding the complex interplay between human genetics, infectious diseases, and evolutionary adaptation.
The C-C chemokine receptor type 5 (CCR5) is a seven-transmembrane, G-protein coupled receptor (GPCR) expressed on the surface of immune cells such as T lymphocytes, macrophages, and dendritic cells [10] [1]. While its primary physiological role involves mediating leukocyte migration and chemotaxis in response to inflammatory cytokines like RANTES (CCL5), MIP-1α (CCL3), and MIP-1β (CCL4), CCR5 gained scientific prominence for its role as a crucial coreceptor for human immunodeficiency virus (HIV) entry [10] [1]. The discovery that HIV-1 utilizes CCR5 alongside the primary CD4 receptor to infect target cells established this receptor as a critical determinant of HIV susceptibility. The viral envelope glycoprotein gp120 first engages CD4, undergoing a conformational change that allows it to subsequently bind to CCR5, ultimately triggering fusion of the viral and cellular membranes [1].
The CCR5Δ32 mutation, a 32-base-pair deletion in the CCR5 gene, produces a truncated protein that fails to localize to the cell surface, thereby conferring resistance to HIV infection in homozygous individuals [31]. This mutation has become a cornerstone for understanding the complex interplay between host genetics, historical selective pressures, and modern therapeutic development. This review examines the evidence linking CCR5Δ32 to historical disease outbreaks, its established role in HIV resistance, and the subsequent translation of this knowledge into novel therapeutic strategies for HIV/AIDS.
Historical Selective Pressures on CCR5Δ32 Distribution
The geographic distribution of the CCR5Δ32 allele is highly non-uniform, with highest frequencies observed in Northern European populations (approximately 10-16% allele frequency) and a pronounced north-to-south gradient declining to near absence in African and Asian populations [32] [31] [33]. This distribution pattern, coupled with the relatively young estimated age of the mutation (between 700 and 3,500 years, though some estimates suggest it may be older), provides strong evidence for intense historical selective pressure rather than genetic drift alone [32] [31].
Table 1: Major Historical Hypotheses for CCR5Δ32 Selection
| Selective Agent | Proposed Mechanism | Supporting Evidence | Contradicting Evidence |
|---|---|---|---|
| Bubonic Plague (Yersinia pestis) | Provided resistance during Black Death pandemics in medieval Europe [31]. | Historical timing correlates with estimated allele age [31]. | Y. pestis infection mechanisms do not clearly involve CCR5; lack of DNA evidence from plague pit victims [32] [31]. |
| Smallpox (Variola major) | conferred survival advantage during frequent smallpox outbreaks [32] [31]. | Smallpox is a strictly human-specific virus with high fatality, creating strong selective pressure; historical records indicate more severe epidemics in N. Europe [32]. | Direct molecular evidence for smallpox virus interaction with CCR5 is limited [31]. |
| Hemorrhagic Fevers (Viral) | Proposed that historical plagues were viral hemorrhagic fevers that utilized CCR5 [31]. | Explains rapid spread and symptoms described in historical accounts [31]. | Lacks direct pathogen identification from historical remains; considered a speculative hypothesis [31]. |
| The Viking Hypothesis | Suggests CCR5Δ32 spread via migration and raids from a Northern European origin [32] [31]. | Correlates with the allele's geographic distribution pattern [32] [31]. | Does not explain the initial selective advantage that increased the allele frequency in the source population [32]. |
Mathematical modeling of the allele's spread supports the conclusion that it underwent strong positive selection. Spatially explicit models indicate that with a selective advantage estimated between 5-35% for heterozygous carriers, combined with dispersal over long distances (>100 km/generation), the current distribution could be achieved within the estimated timeframe [32]. The restriction of the allele to Europe and Western Asia is likely due to insufficient time for wider dispersal rather than a lack of selective advantage elsewhere [32].
CCR5Δ32 Mutation: Molecular Mechanism and HIV Resistance
The molecular basis for HIV resistance stems from the impact of the Δ32 mutation on the CCR5 protein. The 32-bp deletion causes a frameshift and the introduction of a premature stop codon, resulting in a severely truncated and non-functional receptor that is retained intracellularly and degraded [31]. Consequently, the R5-tropic HIV-1 strains, which are predominantly responsible for viral transmission and establishment of infection, cannot effectively bind to and enter host cells [10] [1].
The level of HIV protection is genotype-dependent, as summarized in Table 2.
Table 2: CCR5 Genotypes and Their Correlation with HIV-1 Susceptibility and Disease Progression
| Genotype | Receptor Expression | HIV-1 Susceptibility | Clinical Outcome |
|---|---|---|---|
| CCR5/CCR5 (Wild-type Homozygous) | Normal expression of functional CCR5 [34]. | Fully susceptible to R5-tropic HIV-1 infection [34]. | Standard rate of disease progression [34]. |
| CCR5/CCR5Δ32 (Heterozygous) | Reduced number of functional CCR5 receptors on the cell surface [34] [35]. | Reduced susceptibility to infection; infected individuals have lower pre-AIDS viral loads [34] [35]. | Slower progression to AIDS, delayed by approximately 2 years on average [34]. |
| CCR5Δ32/CCR5Δ32 (Mutant Homozygous) | No functional CCR5 expression on the cell surface [34] [31]. | Highly resistant to infection by R5-tropic HIV-1 strains [34] [31]. | Near-complete protection against HIV infection; rare cases of infection involve X4 or dual-tropic viruses [34] [10]. |
It is critical to note that the protection conferred by the CCR5Δ32/Δ32 genotype is specific for R5-tropic viruses. There are documented cases of HIV infection in individuals with this genotype, typically involving X4-tropic or dual-tropic viral strains that utilize the CXCR4 coreceptor instead [10].
Diagram 1: HIV-1 Entry Pathway. The diagram illustrates the sequential steps of HIV-1 entry, highlighting the critical role of the CCR5 coreceptor. The pathway for the alternative CXCR4 coreceptor, used by X4-tropic viruses, is also shown.
Therapeutic Applications Targeting CCR5
CCR5Δ32-Based Stem Cell Transplantation
The proof-of-concept for curing HIV through CCR5 ablation was established by allogeneic hematopoietic stem cell transplantation (HSCT) from CCR5Δ32/Δ32 donors to HIV-positive patients with hematological malignancies [36]. The "Berlin patient" (2009) and "London patient" (2019) represent landmark cases where this procedure led to long-term HIV remission and functional cure, with no rebound of plasma HIV-1 RNA after cessation of antiretroviral therapy (ART) [36]. A more recent 2023 study reported on a 53-year-old male (IciStem no. 19) who received CCR5Δ32/Δ32 HSCT for acute myeloid leukemia and remained in remission 48 months after analytical treatment interruption, displaying no signs of replication-competent virus despite sporadic traces of HIV DNA [36]. These successes demonstrate that reconstituting the immune system with CCR5-deficient cells can effectively eradicate the viral reservoir, providing a viable path to cure.
CCR5 Inhibitors
The development of CCR5 antagonists represents a direct pharmacological translation of the protective Δ32 mechanism. Maraviroc, approved by the FDA in 2007, is a small molecule that acts as a negative allosteric modulator of CCR5, inducing conformational changes that prevent gp120 binding without triggering signaling pathways [35] [1]. It is indicated for treatment-experienced patients infected with exclusively R5-tropic virus, as determined by tropism assays like the Trofile test [35]. The clinical use of maraviroc underscores the importance of determining viral tropism prior to therapy initiation to avoid treatment failure and potential selection for X4-tropic viruses [35].
Gene Editing Strategies
Advanced gene editing technologies, particularly CRISPR-Cas9, are being harnessed to mimic the CCR5Δ32 mutation in autologous or allogeneic cells [37]. This strategy aims to create a continuous supply of HIV-resistant immune cells, potentially obviating the need for matched CCR5Δ32/Δ32 donors. Research efforts focus on engineering hematopoietic stem cells or induced pluripotent stem cells (iPSCs) to disrupt the CCR5 locus [37]. Accurate quantification of editing efficiency is crucial, and methods like droplet digital PCR (ddPCR) have been developed to detect CCR5Δ32 mutant alleles in heterogeneous cell mixtures with high sensitivity, capable of detecting down to 0.8% of mutant cells [37].
Experimental Toolkit for CCR5 Research
Table 3: Essential Research Reagents and Assays in CCR5 and HIV Research
| Reagent / Assay | Primary Function | Key Features and Applications |
|---|---|---|
| Trofile Tropism Assay (Monogram Biosciences) | Determines HIV-1 coreceptor usage (tropism) in patient plasma samples [35]. | Phenotypic assay; essential prior to CCR5 antagonist therapy; distinguishes R5, X4, and dual/mixed tropic viruses [35]. |
| Droplet Digital PCR (ddPCR) | Absolute quantification of target DNA sequences, including CCR5Δ32 and HIV reservoir metrics [36] [37]. | High precision; used to quantify CCR5Δ32 editing efficiency and trace levels of HIV DNA in cure research [36] [37]. |
| CRISPR-Cas9 System | Genome editing to create targeted knockout of the CCR5 gene in cell lines and primary cells [37]. | Tool for generating the CCR5Δ32 mutation de novo; enables development of autologous cell therapies [37]. |
| Quantitative Viral Outgrowth Assay (qVOA) | Measures the frequency of CD4+ T cells harboring replication-competent HIV [36]. | Gold-standard method for quantifying the latent HIV reservoir in cure trials [36]. |
| Antibodies for CCR5 (e.g., 3A9, HEK/1/85a) | Detect CCR5 surface expression and conformational states via flow cytometry or immunofluorescence [1]. | Used to study receptor density, internalization, and the impact of post-translational modifications (e.g., sulfation) [1]. |
Diagram 2: Gene Therapy Workflow. A generalized workflow for developing an autologous gene therapy for HIV using CRISPR-Cas9 to introduce the CCR5Δ32 mutation into a patient's own cells, such as Hematopoietic Stem Cells (HSCs).
The journey of the CCR5Δ32 allele from a genetic curiosity to a central pillar of HIV therapeutics exemplifies the power of translating evolutionary insights into clinical innovation. The uneven geographic distribution of this mutation provides a compelling, though not yet fully resolved, record of historical selective pressures, potentially from pathogens like smallpox. Its profound resistance to HIV infection unveiled a critical vulnerability of the virus, directly enabling the development of CCR5-targeted therapies, including the small-molecule inhibitor maraviroc and the groundbreaking CCR5Δ32-based stem cell transplants that have produced the only documented cures of HIV infection to date. Future research will continue to refine our understanding of the historical forces that shaped this allele's prevalence while pushing the boundaries of gene editing and cell therapy to make such cures more accessible and less invasive, solidifying CCR5's status as a masterclass in the transition from evolutionary biology to clinical medicine.
The C-C chemokine receptor type 5 (CCR5) serves as a principal coreceptor for human immunodeficiency virus type 1 (HIV-1) entry into target cells. This G protein-coupled receptor, expressed on macrophages, monocytes, and T-cells, facilitates infection by mediating viral attachment and entry in conjunction with CD4 receptor binding [1]. The discovery that a 32-base pair deletion (CCR5Δ32) in the CCR5 gene confers resistance to HIV-1 infection revolutionized understanding of host genetic factors in viral susceptibility [38]. This mutation results in a truncated, non-functional receptor that fails to reach the cell surface, thereby impeding HIV-1 cellular entry [25]. The differential impact of homozygous versus heterozygous CCR5Δ32 genotypes on HIV-1 susceptibility and disease progression represents a critical area of study for understanding viral pathogenesis and developing therapeutic interventions.
Molecular Mechanisms of CCR5Δ32 Mutation
Genetic Basis and Protein Expression
The CCR5Δ32 variant is characterized by a 32-base pair deletion in the coding region of the CCR5 gene on chromosome 3 (3p21.31). This deletion causes a frameshift mutation that introduces a premature stop codon, resulting in a severely truncated and non-functional receptor protein that is retained intracellularly and degraded [25]. In homozygous individuals (Δ32/Δ32), CCR5 surface expression is virtually absent, creating a biological barrier to infection by CCR5-tropic (R5) HIV-1 strains, which constitute the predominantly transmitted variants [25] [38]. In heterozygous individuals (+/Δ32), CCR5 surface expression is reduced approximately 50-70% compared to wild-type homozygotes (+/+) due to haploinsufficiency and possible dominant-negative effects of the mutant protein [39] [40].
Implications for HIV-1 Entry
The structural integrity of CCR5 is essential for HIV-1 envelope glycoprotein gp120 binding following initial CD4 attachment. The V3 loop of gp120 interacts with extracellular domains of CCR5, particularly the N-terminus and second extracellular loop [1]. The Δ32 mutation disrupts this interaction by eliminating functional receptor expression (homozygotes) or reducing available binding sites (heterozygotes). Post-translational modifications of CCR5, including sulfation of tyrosine residues 3, 10, 14, and 15 in the N-terminal region, further modulate HIV-1 entry capability by affecting chemokine and viral envelope binding affinity [1].
Epidemiological Impact on HIV-1 Susceptibility
Genotype Distribution in Global Populations
The CCR5Δ32 allele demonstrates striking geographical variation, with highest prevalence in Northern European populations (allele frequency: 4-16%) and near absence in African, Asian, and Indigenous American populations [39] [25]. This distribution potentially contributes to differential genetic susceptibility to HIV-1 across ethnic groups, alongside behavioral and social factors [39].
Table 1: CCR5Δ32 Genotype Frequencies in HIV-1 Seropositive and Seronegative Cohorts
| Population | Genotype | HIV-1 Seronegative | HIV-1 Seropositive | Odds Ratio (95% CI) |
|---|---|---|---|---|
| Women's Interagency HIV Study (WIHS) [39] | +/+ | 513 | 1940 | Reference |
| +/Δ32 | 45 | 107 | 0.63 (0.44-0.90) | |
| Δ32/Δ32 | 1 | 0 | Undetermined | |
| Meta-Analysis (24 studies) [12] | +/+ | 5,834 | 4,493 | Reference |
| +/Δ32 | 449 | 293 | 1.16 (1.02-1.32) | |
| Δ32/Δ32 | 6 | 12 | 0.25 (0.09-0.68) | |
| High-Risk Caucasian MSM [41] | +/+ | 1,255 | 1,102 | Reference |
| +/Δ32 | 342 | 189 | 0.30 (0.08-0.97) | |
| Δ32/Δ32 | 40 | 1 | <0.05 |
Differential Protection by Genotype
The protective effect of CCR5Δ32 demonstrates clear gene dosage dependence. Homozygous individuals show near-complete resistance to CCR5-tropic HIV-1 transmission, with only rare case reports of infection possibly involving alternative coreceptors (e.g., CXCR4-using variants) [38] [12]. Heterozygous individuals exhibit intermediate protection, with meta-analyses demonstrating significantly reduced infection risk (OR=0.63-0.71) compared to wild-type homozygotes, particularly among high-exposure cohorts [39] [41] [12]. This partial protection stems from reduced CCR5 surface density, which decreases viral entry efficiency and initial propagation within mucosal tissues following exposure.
Disease Progression in Infected Individuals
Clinical Outcomes by Genotype
Among HIV-1-infected individuals, CCR5Δ32 genotype significantly influences disease progression kinetics. Heterozygous carriers typically experience delayed progression to AIDS by 2-4 years compared to wild-type homozygotes, with slower CD4+ T-cell decline and lower viral loads during asymptomatic stages [34] [38] [41]. This delayed progression reflects reduced CCR5-mediated viral replication and possibly enhanced antiviral immune responses due to preserved CCR5-mediated chemotaxis at reduced receptor density.
Table 2: Disease Progression Parameters by CCR5 Genotype in HIV-1 Infected Individuals
| Parameter | Wild-type (+/+) | Heterozygous (+/Δ32) | Homozygous (Δ32/Δ32) |
|---|---|---|---|
| Surface CCR5 Expression | 100% | 30-50% | <1% |
| HIV-1 Susceptibility | High | Intermediate | Highly resistant |
| Progression to AIDS | 7-10 years | 9-12 years | Extremely rare |
| Pre-AIDS Viral Load | High | Intermediate | Not applicable |
| CD4+ Decline Rate | Rapid | Slowed | Not applicable |
CCR5 Promoter Polymorphisms and Modifying Effects
Beyond the Δ32 variant, single nucleotide polymorphisms (SNPs) in the CCR5 promoter region significantly modify HIV-1 disease progression by regulating transcriptional activity and receptor density [42] [40]. Specific haplotypes (HHA-HHG*2) demonstrate varying promoter activities that correlate with CCR5 expression levels and disease course. The HHC haplotype is associated with slower CD4+ decline and delayed AIDS onset, while the HHE haplotype correlates with accelerated disease, particularly in heterozygous individuals where the non-Δ32 haplotype determines phenotypic expression [42] [40]. This haplotype-phenotype relationship explains heterogeneity in clinical outcomes among Δ32 heterozygotes and underscores the importance of complete CCR5 genotyping in prognostic assessments.
Experimental Approaches and Methodologies
CCR5 Genotyping Protocols
DNA Extraction and Quality Control
- Source: Peripheral blood mononuclear cells (PBMCs) or buccal swabs
- Method: Phenol-chloroform extraction or commercial silica-membrane kits
- Quality Assessment: Spectrophotometric (A260/A280 ratio 1.8-2.0) and gel electrophoresis
PCR Amplification of CCR5 Locus
- Primers: Forward: 5'-CTCACACCCTGTGCCTCTT-3', Reverse: 5'-TCATTTCGACACCGAAGCAG-3'
- Reaction Mix: 1X PCR buffer, 1.5mM MgCl₂, 200μM dNTPs, 0.2μM each primer, 1.25U Taq polymerase, 100ng genomic DNA
- Cycling Conditions: Initial denaturation 95°C/5min; 35 cycles of 95°C/30s, 60°C/30s, 72°C/45s; final extension 72°C/7min
Genotype Analysis
- Amplicon Sizes: Wild-type: 332bp, Δ32: 300bp
- Detection: 2-3% agarose gel electrophoresis with ethidium bromide staining
- Validation: Restriction fragment length polymorphism (RFLP) or direct Sanger sequencing [39] [40]
Promoter Activity Assays
Luciferase Reporter Constructs
- Vector: pGL3-Basic or similar promoterless luciferase vectors
- Insert: CCR5 promoter haplotypes (HHA-HHG) cloned upstream of luc gene
- Transfection: HEK293T or Jurkat cells using lipid-based transfection reagents
Relative Promoter Activity (RPA) Measurement
- Normalization: Co-transfection with Renilla luciferase (pRL vectors)
- Assay: Dual-luciferase reporter system
- Calculation: Firefly/Renilla luciferase ratio normalized to HHA (ancestral) haplotype
- Combined RPA (CRPA): Sum of both allele RPAs for correlation with clinical parameters [40]
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for CCR5-HIV Studies
| Reagent/Category | Specific Examples | Research Application |
|---|---|---|
| Genotyping Reagents | CCR5-specific primers, restriction enzymes (for RFLP), Taq polymerase | CCR5Δ32 and promoter haplotype determination |
| Cell Lines | PM1, CEMx174, HEK293T, TZM-bl | Viral replication studies, promoter activity assays, infection efficiency |
| Antibodies | Anti-CCR5 (clone 2D7/3A9), anti-CD4, anti-CXCR4 | Flow cytometry, receptor quantification, blocking studies |
| CCR5 Antagonists | Maraviroc, Vicriviroc, Aplaviroc | Therapeutic mechanism studies, receptor blockade experiments |
| Cytokines/Chemokines | RANTES/CCL5, MIP-1α/CCL3, MIP-1β/CCL4 | Ligand competition assays, receptor internalization studies |
| Viral Strains | CCR5-tropic (Bal, JR-FL), CXCR4-tropic (NL4-3), dual-tropic | Tropism studies, coreceptor usage assays, neutralization tests |
Therapeutic Implications and Drug Development
CCR5-Targeted Therapeutics
The CCR5Δ32 natural experiment validated CCR5 inhibition as an antiretroviral strategy, leading to development of CCR5 antagonists. Maraviroc, the first FDA-approved CCR5 antagonist, acts as an allosteric inhibitor that stabilizes CCR5 in an inactive conformation incapable of supporting HIV-1 entry [25] [1]. Its clinical efficacy demonstrates that partial receptor blockade (mimicking heterozygous phenotype) provides substantial antiviral effect, supporting the concept that complete receptor inhibition is unnecessary for therapeutic benefit [41]. Treatment strategies based on CCR5 genotyping may optimize antiretroviral regimens, particularly for patients with limited therapeutic options or drug resistance.
Gene Editing Approaches
Novel therapeutic approaches include CCR5 gene editing using CRISPR/Cas9 or zinc finger nucleases to mimic the Δ32 phenotype in hematopoietic stem cells or CD4+ T-cells. The "Berlin Patient" and "London Patient" who achieved HIV-1 remission following CCR5Δ32/Δ32 stem cell transplantation for hematological malignancies provide proof-of-concept for CCR5 disruption as a curative strategy [25]. Current efforts focus on enhancing editing efficiency and safety profiles for clinical application.
The differential impact of homozygous and heterozygous CCR5Δ32 genotypes on HIV-1 susceptibility and disease progression provides a powerful model for understanding host-pathogen interactions. The gene dosage effect observed—with homozygotes enjoying near-complete protection and heterozygotes showing partial protection and slowed disease progression—has informed therapeutic development and underscored the importance of host genetics in infectious disease outcomes. Future research directions include elucidating how CCR5 heterogeneity influences tissue-specific viral reservoirs, understanding compensatory mechanisms in CCR5-deficient individuals, and developing next-generation therapies that exploit CCR5 biology while minimizing off-target effects. The continued study of these natural genetic variants will undoubtedly yield further insights into HIV-1 pathogenesis and treatment.
Harnessing CCR5 Biology: From Gene Editing to Clinical HIV Cure Strategies
The C-C chemokine receptor type 5 (CCR5) is a class A G protein-coupled receptor (GPCR) that plays a critical role in the human immune system, primarily mediating the migration of monocytes and T-helper 1 (Th1) cells to sites of inflammation [43] [10]. Its path to prominence began in the 1990s with its identification as a crucial coreceptor for human immunodeficiency virus type 1 (HIV-1) entry into target cells [43] [1]. HIV-1 infection is a two-step process that initially involves the binding of the viral envelope glycoprotein gp120 to the CD4 receptor on the host cell surface, followed by a conformational change that allows gp120 to interact with a coreceptor—predominantly CCR5 or CXCR4 [1] [44]. This second interaction defines viral tropism; viruses that utilize CCR5 are designated R5-tropic and are particularly significant as they are predominantly involved in transmission events and are the predominant strain during the early and chronic phases of HIV-1 infection [45] [44].
The profound importance of CCR5 in HIV-1 pathogenesis was unequivocally demonstrated by the discovery of the CCR5Δ32 genetic variant. This natural 32-base pair deletion in the CCR5 gene results in a truncated protein that is not expressed on the cell surface [10] [45]. Individuals who are homozygous for the CCR5Δ32 allele are almost completely resistant to HIV-1 infection, while heterozygotes exhibit slower disease progression and lower viral loads [34] [44]. This natural resistance provided a powerful genetic validation of CCR5 as a therapeutic target, spurring the development of entry inhibitors designed to pharmacologically mimic this protective effect. This whitepaper explores the development, mechanism, and application of CCR5 blockade, with a focus on maraviroc, situating this therapeutic strategy within the broader context of CCR5 biology and Δ32 mutation research.
The Biological Foundation of CCR5
Structural Biology and Physiological Function
CCR5 is a member of the GPCR superfamily, characterized by a structure of seven transmembrane α-helices connected by three extracellular and three intracellular loops [10]. Its N-terminal domain is extracellular and is subject to critical post-translational modifications, including sulfation of tyrosine residues, which enhance the binding of chemokines and the HIV-1 gp120 protein [1]. The receptor is expressed on a range of immune cells, including activated T lymphocytes, macrophages, dendritic cells, and microglia in the central nervous system [10] [44].
In its physiological role, CCR5 is activated by its endogenous chemokine ligands, such as CCL3 (MIP-1α), CCL4 (MIP-1β), and CCL5 (RANTES) [10]. Upon ligand binding, CCR5 initiates intracellular signaling cascades primarily through associated G-proteins. This triggers processes like phospholipase C activation and inositol-triphosphate (IP3) generation, leading to cytosolic calcium release and actin polymerization, which drives immune cell chemotaxis towards the source of the chemokine gradient [10]. Beyond chemotaxis, CCR5 also functions in immune synapses, acting as a costimulatory molecule that can enhance T-cell proliferation and cytokine secretion in response to antigens [10].
The CCR5Δ32 Mutation: Population Genetics and Selective Pressure
The CCR5Δ32 allele is found predominantly in populations of European descent, with allelic frequencies ranging from 0 to as high as 0.29 in some groups, but it is almost absent in African and Asian populations [34]. Mathematical modeling of heterosexual HIV epidemics has shown that the presence of the CCR5Δ32 allele in a population can lower the overall prevalence of HIV/AIDS compared to populations lacking the allele [34]. Furthermore, these models indicate that HIV infection itself exerts a selective pressure, leading to a gradual increase in the frequency of the CCR5Δ32 allele over time in affected populations [34].
The landmark case of an HIV-positive patient with acute myeloid lymphoma who received a transplant of CCR5Δ32/Δ32 homozygous stem cells and experienced long-term control of HIV infection without antiretroviral therapy further underscores the therapeutic potential of targeting this receptor [10]. This case, along with the natural resistance of Δ32 homozygotes, confirmed that CCR5 is the major portal of entry for HIV and that its blockade or absence does not cause severe immunodeficiency, making it a safe and viable target for drug development.
Table 1: Impact of CCR5Δ32 Genotype on HIV-1 Infection
| Genotype | CCR5 Expression | Susceptibility to R5-tropic HIV-1 | Clinical Outcome |
|---|---|---|---|
| CCR5/CCR5 (Wild-type homozygote) | Normal | High | Standard rate of disease progression |
| CCR5/CCR5Δ32 (Heterozygote) | Reduced | Moderate | Delayed progression to AIDS, lower viral loads [34] [44] |
| CCR5Δ32/CCR5Δ32 (Mutant homozygote) | Not expressed [45] | Highly resistant [45] | Near-complete protection from infection [34] [44] |
The Development of CCR5 Antagonists
Maraviroc: The First-in-Class CCR5 Antagonist
Maraviroc (brand name Selzentry) is a small-molecule CCR5 antagonist developed by Pfizer and approved by the US FDA in 2007 for use in combination with other antiretroviral agents for treatment-experienced adults infected with only CCR5-tropic HIV-1 [46] [45]. It is the first and, to date, the only licensed oral drug in its class.
Mechanism of Action
Maraviroc functions as an allosteric, non-competitive inhibitor of the CCR5 receptor [44]. It binds to a hydrophobic pocket formed by the transmembrane helices of CCR5, inducing a conformational change in the receptor's extracellular domains [45]. This structural alteration does not prevent the natural chemokines from binding but makes the receptor unrecognizable to the V3 loop of the HIV-1 gp120 protein, thereby blocking the essential coreceptor step in viral entry [45] [44]. It is important to note that maraviroc is a tropism-dependent therapeutic; it is only effective against R5-tropic viruses and has no efficacy against viruses that use the CXCR4 coreceptor (X4-tropic or dual-tropic viruses) [45].
Clinical Efficacy and Safety
The efficacy and safety of maraviroc were established in the MOTIVATE 1 and 2 trials. These were double-blind, placebo-controlled, multicenter phase 2b/3 studies conducted in treatment-experienced patients with R5-tropic virus. Patients received maraviroc once or twice daily plus an optimized background therapy (OBT), compared to placebo plus OBT.
Table 2: Key Efficacy Outcomes from MOTIVATE 1 & 2 Trials (Pooled Data at 48 Weeks) [45]
| Treatment Group | Mean Change in HIV-1 RNA (log10 copies/mL) | Mean CD4+ Cell Count Increase (cells/μL) |
|---|---|---|
| Maraviroc Once Daily + OBT | -1.66 to -1.72 | +113 to +122 |
| Maraviroc Twice Daily + OBT | -1.82 to -1.87 | +122 to +128 |
| Placebo + OBT | -0.76 to -0.80 | +54 to +69 |
The trials demonstrated that maraviroc was generally well-tolerated. The most common adverse events were cough, fever, upper respiratory tract infections, rash, and musculoskeletal symptoms. A theoretical concern with CCR5 blockade was the potential promotion of X4-tropic viruses, which are associated with more rapid disease progression. However, accumulated clinical data has not substantiated this fear [44].
Other CCR5-Targeting Agents
While maraviroc is the only approved small-molecule CCR5 antagonist, other agents have been developed. Vicriviroc reached Phase III clinical trials but has not been approved, and aplaviroc development was halted due to hepatotoxicity [44]. Additionally, biological agents have been explored, including:
- PRO 140: A humanized monoclonal antibody that binds to the extracellular domain of CCR5, blocking gp120 association. It has demonstrated potent antiviral activity in clinical trials [44].
- HGS004: Another human monoclonal antibody against CCR5 that has shown in vivo activity [44].
Experimental and Clinical Methodologies
Determining Viral Tropism
The effective use of maraviroc is contingent upon the confirmed presence of an R5-tropic virus. Two primary methodologies are used for tropism testing, each with distinct advantages and limitations.
Table 3: Comparison of Tropism Testing Methodologies
| Method | Principle | Procedure | Advantages | Disadvantages |
|---|---|---|---|---|
| Phenotypic Assay (e.g., Trofile) | Uses cell culture to determine which coreceptor the virus can use for entry [45]. | Patient-derived virus pseudovirions are engineered to contain a reporter gene (e.g., luciferase). They are used to infect cell lines expressing either CCR5 or CXCR4. Infection success is measured by reporter signal [45]. | Considered the historical gold standard; directly measures function. | Technically complex, expensive (~$750-$1000), time-consuming (up to 4 weeks), requires specialized labs [45]. |
| Genotypic Assay | Analyzes the nucleotide sequence of the HIV-1 V3 loop region of the env gene, the primary determinant of coreceptor usage [45] [44]. | Viral RNA is extracted from patient plasma. The V3 loop region is amplified by RT-PCR and sequenced. The sequence is analyzed using bioinformatic algorithms (e.g., geno2pheno[coreceptor], WebPSSM) to predict tropism [45]. | Faster (days), less expensive, more accessible. | Potentially lower sensitivity/specificity than phenotypic assays; performance varies with the algorithm used [45]. |
The following diagram illustrates the workflow for determining the appropriate use of maraviroc based on tropism testing:
The Scientist's Toolkit: Key Research Reagents
Research into CCR5 biology and the development of entry inhibitors relies on a suite of specialized reagents and tools.
Table 4: Essential Research Reagents for CCR5 Investigation
| Reagent / Tool | Function and Application | Key Details |
|---|---|---|
| CCR5-Specific Monoclonal Antibodies | Used for flow cytometry (FACS) to quantify CCR5 surface expression on different cell types [1]; some can block HIV-1 entry (e.g., PRO 140) [44]. | Examples: mAb 3A9 (binds sulfated N-terminal peptide), HEK/1/85a [1]. |
| Recombinant Chemokines | Natural ligands (CCL3, CCL4, CCL5) used in competition assays to study HIV-1 entry inhibition and CCR5 signaling pathways [10]. | CCL3, CCL4, and CCL5 display HIV-inhibiting properties in vitro [10]. |
| CCR5Δ32 Genotyping Assays | To identify the 32-bp deletion in the CCR5 gene for population genetics studies or correlating genotype with disease progression. | PCR-based methods using specific primers flanking the deletion region. |
| Phenotypic Tropism Assays | To definitively characterize the coreceptor usage (tropism) of patient-derived HIV-1 strains in a clinical or research setting [45]. | Commercial version: Trofile assay [45]. |
| Genotypic Tropism Algorithms | Bioinformatics tools to predict HIV-1 coreceptor usage from the V3 loop sequence [45]. | Examples: geno2pheno[coreceptor], WebPSSM, Webcat [45]. |
| CCR5-Expressing Cell Lines | Engineered cell lines (e.g., HEK-293, U87) stably expressing CD4 and CCR5 for in vitro viral infection and neutralization assays. | Essential for studying viral entry and testing entry inhibitors. |
Beyond HIV: Therapeutic Repurposing of CCR5 Blockade
The role of CCR5 extends beyond HIV infection, with research implicating it in other viral infections, autoimmune diseases, and cancer metastasis [28] [47]. Notably, CCR5 signaling is involved in neuroinflammation. The receptor becomes increasingly active in the aging brain, contributing to synaptic disruption and cognitive decline [48]. This has spurred investigations into repurposing maraviroc for Alzheimer's disease. Researchers are developing nanoparticle formulations of maraviroc to enhance its delivery across the blood-brain barrier, aiming to reduce neuroinflammation and slow disease progression with potentially fewer side effects than current anti-amyloid therapies [48].
Furthermore, CCR5 antagonists are being explored for use in organ transplantation to prevent rejection and as part of combination therapies with other entry inhibitors or agents like rapamycin that downregulate CCR5 expression [44]. However, the broad use of CCR5 blockade requires caution, as the CCR5Δ32 mutation has been associated with increased susceptibility and worse outcomes in certain viral infections, such as West Nile virus [28] [44].
The journey from the discovery of CCR5 as an HIV-1 coreceptor and the protective CCR5Δ32 mutation to the development and clinical implementation of maraviroc represents a triumph of translational medicine. Maraviroc, as the first-in-class CCR5 antagonist, validated a novel host-targeted strategy against HIV-1, offering a potent option for patients with R5-tropic virus. Its use necessitates rigorous tropism testing, for which both phenotypic and genotypic methods have been refined. The experimental tools developed to study CCR5 continue to reveal its multifaceted roles in human physiology and disease. As research progresses, the therapeutic application of CCR5 blockade is poised to expand beyond HIV, offering new avenues for treating neuroinflammatory and other immune-mediated disorders, while reminding us of the importance of understanding the complex and context-dependent functions of immune receptors.
The C-C chemokine receptor type 5 (CCR5) serves as a principal co-receptor for human immunodeficiency virus (HIV) entry into CD4+ T-cells, making it a critical therapeutic target. The discovery that a natural 32-base pair deletion in the CCR5 gene (CCR5Δ32) confers resistance to HIV-1 infection in homozygous individuals provided a genetic blueprint for curing HIV [49] [50]. This breakthrough, exemplified by the "Berlin" and "London" patients who were cured of HIV after receiving CCR5Δ32/Δ32 stem cell transplants, demonstrated the profound therapeutic potential of CCR5 disruption [3]. However, the scarcity of naturally compatible CCR5Δ32 donors has driven the development of programmable gene-editing technologies to artificially disrupt CCR5 in patient cells. This review provides a comparative analysis of the three major gene-editing platforms—Zinc Finger Nucleases (ZFNs), Transcription Activator-Like Effector Nucleases (TALENs), and CRISPR/Cas9—within the context of CCR5-targeted HIV therapy, examining their mechanisms, efficiencies, and clinical applications.
Technological Mechanisms and Evolution
Fundamental Mechanisms of Programmable Nucleases
The three major gene-editing platforms, while distinct in architecture, share a common functional principle: creating double-strand breaks (DSBs) in genomic DNA at predetermined sites, which are then repaired by the cell's endogenous repair machinery.
Zinc Finger Nucleases (ZFNs) are synthetic fusions of a zinc finger DNA-binding domain to the FokI cleavage domain. Each zinc finger typically recognizes a 3-base pair DNA triplet, and arrays of 3-6 fingers are assembled to confer sequence specificity [51] [52]. The FokI nuclease must dimerize to become active, necessitating the design of two ZFN proteins that bind to opposite DNA strands with appropriate spacing and orientation [51].
Transcription Activator-Like Effector Nucleases (TALENs) similarly fuse a DNA-binding domain to the FokI nuclease. However, TALENs utilize TALE repeat domains derived from plant-pathogenic bacteria, where each repeat recognizes a single nucleotide [51] [52]. This one-to-one recognition code simplifies design and enhances targeting range. Like ZFNs, TALENs require paired binding for FokI dimerization and DSB formation [51].
CRISPR/Cas9 employs a fundamentally different mechanism based on RNA-DNA recognition. The system consists of two components: a Cas9 nuclease and a single guide RNA (sgRNA) that combines the functions of CRISPR RNA (crRNA) and trans-activating crRNA (tracrRNA) [53] [51]. The sgRNA directs Cas9 to complementary genomic sequences adjacent to a Protospacer Adjacent Motif (PAM), typically 5'-NGG-3' for standard Streptococcus pyogenes Cas9 [51]. Cas9 then induces a DSB without requiring dimerization.
Comparative Analysis of Editing Platforms
Table 1: Comprehensive Comparison of Gene-Editing Technologies
| Feature | ZFNs | TALENs | CRISPR/Cas9 |
|---|---|---|---|
| Recognition Mechanism | Protein-DNA (3 bp/finger) | Protein-DNA (1 bp/repeat) | RNA-DNA (20 bp guide + PAM) |
| Cleavage Mechanism | FokI dimerization | FokI dimerization | Cas9 nuclease |
| Target Length | 9-18 bp | 30-40 bp | 22 bp + PAM |
| Design Complexity | Challenging; context-dependent effects | Moderate; modular assembly | Simple; sgRNA synthesis |
| Development Timeline | Weeks-months | Weeks | Days |
| Multiplexing Capacity | Limited | Limited | High (multiple sgRNAs) |
| Primary Advantages | Established clinical experience; smaller size | High specificity; flexible targeting | Easy design; cost-effective; highly versatile |
| Primary Limitations | High cost; difficult design; potential cytotoxicity | Large size challenges delivery; time-consuming cloning | PAM dependency; off-target concerns |
Quantitative Performance and Specificity Analysis
Efficiency and Specificity Benchmarks
Direct comparative studies reveal significant differences in editing performance across platforms. A comprehensive 2021 study using genome-wide unbiased identification of double-stranded breaks enabled by sequencing (GUIDE-seq) to assess off-target activities of ZFNs, TALENs, and SpCas9 targeting human papillomavirus 16 (HPV16) genes found that SpCas9 was more efficient and specific than both ZFNs and TALENs [54]. The study documented dramatically different off-target counts: at the URR gene target, SpCas9 had zero off-targets, TALENs had one off-target, while ZFNs generated 287 off-target sites [54]. Similar patterns emerged at E6 and E7 genes, establishing CRISPR/Cas9's superior specificity profile in this direct comparison.
Table 2: Quantitative Performance Metrics in Comparative Studies
| Metric | ZFNs | TALENs | CRISPR/Cas9 |
|---|---|---|---|
| On-target Efficiency | Variable (dependent on design) | High with optimized designs | Consistently high across targets |
| Off-target Events (URR gene) | 287 | 1 | 0 |
| Off-target Events (E6 gene) | Not reported | 7 | 0 |
| Off-target Events (E7 gene) | Not reported | 36 | 4 |
| Clinical Trial Prevalence | 13 trials (as of 2020) | 6 trials (as of 2020) | 42 trials (as of 2020) |
| Targeting Flexibility | Limited by finger availability | High (simple code) | Very high (easily programmable) |
Clinical Translation and Applications
The progression of these technologies to clinical application demonstrates their therapeutic potential. As of October 2020, there were 13 clinical trials involving ZFNs, 6 for TALENs, and 42 for CRISPR/Cas9 registered on ClinicalTrials.gov [54]. ZFN-mediated CCR5 disruption in CD4+ T cells (SB-728-T) has entered multiple Phase I clinical trials for HIV, demonstrating safety and efficacy in reducing viral DNA copies in treated patients [54] [3]. Similarly, TALEN-edited universal chimeric antigen receptor T cells (UCART19) have induced molecular remission in pediatric B-cell acute lymphoblastic leukemia [54]. The first clinical trial applying CRISPR/Cas9 to CCR5 editing in hematopoietic stem cells (NCT03164135) demonstrated successful engraftment and persistence of edited cells for over 19 months without adverse events [53] [3].
CCR5-Targeted Gene Editing for HIV Therapy
From Natural Mutation to Therapeutic Strategy
The CCR5Δ32 mutation occurs naturally in approximately 1% of Caucasian populations, providing complete resistance to R5-tropic HIV-1 strains when homozygous [53] [49] [50]. This discovery, stemming from population genetics studies in the 1990s, revealed that CCR5 deficiency is compatible with normal health, making it an ideal therapeutic target [49]. Gene editing technologies aim to mimic this natural resistance by creating targeted disruptions in the CCR5 gene, potentially leading to a functional cure for HIV infection.
Current approaches include:
- Ex vivo editing of CD4+ T cells or hematopoietic stem/progenitor cells (HSPCs) followed by reinfusion, creating a protected immune cell population [3]
- Combinatorial strategies targeting both CCR5 and additional antiviral targets, such as the CXCR4 co-receptor or HIV fusion inhibitors [53]
- Multiplexed editing to target viral reservoirs by disrupting HIV long terminal repeat (LTR) regions while protecting host cells via CCR5/CXCR4 ablation [3]
Experimental Protocol: CRISPR/Cas9-Mediated CCR5 Knockout
The following detailed methodology for CCR5 knockout in hematopoietic cells has been successfully employed in clinical trials (NCT03164135) and recent research [53]:
1. sgRNA Design and Preparation:
- Design two sgRNAs targeting the first exon of human CCR5 gene, corresponding to the Δ32 mutation site
- Select sgRNAs with high on-target efficiency and minimal predicted off-target effects using computational tools
- Synthesize sgRNAs using in vitro transcription or commercial synthesis
2. Ribonucleoprotein (RNP) Complex Formation:
- Complex purified Cas9 protein with sgRNAs at optimized ratios (e.g., 6-10μg Cas9 with 2-4μg of each sgRNA)
- Incubate for 10-20 minutes at room temperature to allow RNP assembly
3. Cell Nucleofection:
- Use MT4CCR5 cell line or primary human CD34+ HSPCs as target cells
- Resuspend 1×10^6 cells in nucleofection solution
- Mix cell suspension with RNP complexes and transfer to nucleofection cuvette
- Perform nucleofection using optimized program (e.g., U-014 for HSPCs)
4. Post-nucleofection Analysis:
- Assess editing efficiency at 72 hours post-nucleofection using T7 endonuclease I (T7E1) assay or tracking of indels by decomposition (TIDE)
- Evaluate CCR5 protein expression reduction by flow cytometry and Western blotting
- Determine cell viability using 7-aminoactinomycin D (7AAD) staining
5. Functional Validation:
- Challenge edited cells with R5-tropic HIV-1 strains (e.g., BaL)
- Measure viral replication by p24 ELISA or RT-PCR
- Quantify cell protection via viability assays and CD4 expression maintenance
Combinatorial Approaches for Enhanced Protection
While CCR5 disruption provides resistance to R5-tropic HIV, it offers no protection against X4-tropic strains that emerge in approximately 50% of patients during late-stage infection [53]. To address this limitation, researchers have developed combinatorial strategies:
- Dual CCR5/CXCR4 knockout prevents coreceptor switching by simultaneously disrupting both major HIV co-receptors [3]
- CCR5 knockout plus C46 fusion inhibitor expression provides broad-spectrum protection against both R5 and X4 tropic HIV-1 [53]
- Multiplexed host and viral targeting combining CCR5 ablation with HIV LTR disruption to target latent reservoirs [3]
A 2024 study demonstrated that combining CRISPR/Cas9-mediated CCR5 knockout with lentiviral delivery of the C46 HIV-1 fusion inhibitor provided superior protection against both R5 and X4 tropic HIV-1 compared to single-modality approaches [53]. This synergistic strategy represents the next generation of gene-editing applications for HIV cure research.
Essential Research Reagents and Methodologies
Table 3: Key Research Reagent Solutions for CCR5 Gene Editing Studies
| Reagent Category | Specific Examples | Function/Application |
|---|---|---|
| Editing Platforms | ZFN pairs (CCR5-specific), TALEN pairs (CCR5-specific), SpCas9 + sgRNA constructs | Core editing machinery for CCR5 disruption |
| Delivery Systems | Electroporation/nucleofection devices, Lentiviral vectors, AAV vectors | Introduction of editing components into target cells |
| Validation Assays | T7 Endonuclease I (T7E1) assay, Tracking of Indels by Decomposition (TIDE), GUIDE-seq | Assessment of editing efficiency and specificity |
| Cell Culture Models | MT4CCR5 cell line, Primary human CD34+ HSPCs, Patient-derived CD4+ T cells | Cellular substrates for editing experiments |
| Analysis Tools | Flow cytometry antibodies (anti-CCR5, anti-CD4), HIV p24 ELISA kits, Western blot reagents | Functional validation of editing outcomes |
| Specialized Reagents | Recombinant Cas9 protein, Chemically synthesized sgRNAs, FokI nuclease variants | Components for RNP complex formation |
The evolution of gene-editing technologies from ZFNs to TALENs and CRISPR/Cas9 has progressively enhanced our ability to target the CCR5 receptor for HIV therapy. While ZFNs established the clinical proof-of-concept and TALENs offered improved specificity, CRISPR/Cas9 has emerged as the most versatile and accessible platform due to its simplicity of design, high efficiency, and capacity for multiplexed editing [54] [51] [52]. The successful application of CRISPR/Cas9 in clinical trials for CCR5 editing represents a landmark achievement in the field [53] [3].
Future directions include:
- Enhanced specificity systems such as high-fidelity Cas9 variants and base editors to minimize off-target effects [3] [52]
- Combinatorial targeting strategies addressing both host factors (CCR5/CXCR4) and viral reservoirs for comprehensive HIV eradication [53] [3]
- Improved delivery methodologies including nanoparticle-based systems for in vivo editing [52]
- Synergistic integration with immunotherapy approaches such as CAR-T cells and immune checkpoint modulation [3]
The continued refinement of these gene-editing platforms promises to overcome current limitations in HIV treatment, moving closer to a functional cure for this global health challenge.
Allogeneic hematopoietic stem-cell transplantation (allo-HSCT) from donors with a homozygous CCR5Δ32 mutation has led to the only documented cases of sustained HIV-1 remission, representing a pivotal milestone in the quest for an HIV cure. This whitepaper provides a technical analysis of the seminal "Berlin" and "London" patient cases, framing them within the broader context of CCR5 as an HIV coreceptor and CCR5Δ32 mutation research. We detail the experimental methodologies used to validate HIV cure, summarize quantitative data in structured tables, and explore the mechanistic insights these cases provide. Furthermore, we discuss the translation of these findings into next-generation therapeutic strategies, including gene editing and immunotherapies, which aim to replicate the curative phenotype without the risks of allo-HSCT.
The C-C chemokine receptor type 5 (CCR5) is a seven-transmembrane, G protein-coupled receptor expressed on the surface of various immune cells, including CD4+ T lymphocytes, macrophages, and dendritic cells [11]. Its primary physiological role is to direct immune cell migration (chemotaxis) to sites of inflammation along a gradient of its natural ligands, such as CCL3, CCL4, and CCL5 [11]. For human immunodeficiency virus (HIV-1), CCR5 serves as the primary coreceptor for viral entry, particularly for strains (R5-tropic) that are dominant during initial and chronic phases of infection. Viral entry requires the envelope glycoprotein gp120 to bind sequentially to the CD4 receptor and then to the CCR5 coreceptor, triggering fusion with the host cell membrane [3] [11].
The CCR5Δ32 genetic variant is a 32-base-pair deletion in the coding region of the CCR5 gene. This mutation results in a frameshift and the production of a truncated, non-functional protein that is not expressed on the cell surface [11]. Individuals who are homozygous for CCR5Δ32 (genotype CCR5Δ32/Δ32) lack functional CCR5 receptors on their CD4+ T cells and exhibit a high degree of resistance to infection with R5-tropic HIV-1 [34] [11]. The global distribution of this allele is uneven, with the highest frequencies (averaging ~10%) found in populations of Northern European descent, and it is virtually absent in African, Asian, and Native American populations [34] [11]. This natural resistance mechanism provided the foundational rationale for targeting CCR5 in curative strategies.
Case Analysis: Berlin and London Patients
The Berlin Patient (Timothy Ray Brown)
The Berlin Patient was the first documented case of HIV-1 cure. He underwent two allo-HSCT procedures in 2006 and 2007 to treat acute myeloid leukemia (AML) [55] [56]. His donor was homozygous for the CCR5Δ32 mutation. The transplant conditioning regimen included total body irradiation (TBI) and chemotherapy, and the patient experienced significant complications, including graft-versus-host disease (GvHD) [57]. Despite these challenges, he discontinued antiretroviral therapy (ART) at the time of his first transplant and remained aviremic until his passing in 2020 from recurrent leukemia [57].
The London Patient (Participant 36 in the IciStem Cohort)
The London Patient underwent a single allo-HSCT in 2016 for Hodgkin's Lymphoma using stem cells from a CCR5Δ32/Δ32 donor [55] [58]. His conditioning regimen involved LACE chemotherapy (Lomustine, Cyclophosphamide, Ara-C, Etoposide) and Alemtuzumab (anti-CD52) for in vivo T-cell depletion, but did not include total body irradiation [55] [56]. This demonstrated that a less aggressive, reduced-intensity conditioning could still achieve remission. The patient experienced only mild gut GvHD. ART was interrupted 16 months post-transplant, and HIV-1 remission has been maintained for over 30 months post-treatment interruption, with extensive testing confirming the absence of replication-competent virus [58].
Table 1: Patient Demographics and Clinical History
| Parameter | Berlin Patient | London Patient |
|---|---|---|
| Year of Transplant | 2006 & 2007 | 2016 |
| Condition Treated | Acute Myeloid Leukemia | Hodgkin's Lymphoma |
| CCR5 Genotype (Pre-HSCT) | CCR5wt/wt | CCR5wt/wt |
| Donor CCR5 Genotype | CCR5Δ32/Δ32 | CCR5Δ32/Δ32 |
| ART Interruption | At first transplant | 16 months post-transplant |
Table 2: Transplant Procedure and Outcome
| Parameter | Berlin Patient | London Patient |
|---|---|---|
| Conditioning Regimen | Total Body Irradiation + Chemotherapy | LACE Chemotherapy + Alemtuzumab (No Irradiation) |
| GvHD | Severe | Mild (gut, Grade 1) |
| Chimerism | Full donor | Full donor (99% in T-cells) |
| Time off ART | ~13 years | >30 months (as of 2020) |
| HIV Status | Cured | Cured |
Experimental Protocols for Validating HIV Cure
Rigorous and multi-faceted experimental approaches are required to confirm HIV-1 remission or cure. The following protocols were employed in the London Patient case and represent the gold standard [55] [58].
Reservoir Quantification and Viral Load Assays
- Ultra-Sensitive Viral Load Testing: Plasma HIV-1 RNA levels were measured using an assay with a detection limit of <1 copy per mL [55] [58]. Testing was performed weekly for the first three months post-ART interruption and monthly thereafter.
- Total Cell-Associated HIV-1 DNA PCR: Total HIV-1 DNA in peripheral blood mononuclear cells (PBMCs) and purified CD4+ T cells was quantified using droplet digital PCR (ddPCR). Targets included the HIV-1 long-terminal repeat (LTR) and Gag genes, with results reported as copies per million cells [55] [58].
- Intact Proviral DNA Assay (IPDA): A multiplex ddPCR assay targeting the packaging signal (ψ) and envelope (env) regions was used to distinguish between intact and defective proviruses [58].
Viral Outgrowth and Replication Competence
- Quantitative Viral Outgrowth Assay (QVOA): Resting CD4+ T cells are isolated and maximally stimulated to induce latent virus. The culture supernatants are then tested for p24 antigen or viral RNA to detect replication-competent virus. Results are reported in Infectious Units Per Million (IUPM) resting CD4+ T cells. In the London Patient, a total of 24 million resting CD4+ T cells were assayed across three time points with no reactivatable virus, yielding an IUPM of <0.029 [55] [58].
Viral Tropism and Cell Susceptibility
- Coreceptor Usage Prediction: The HIV-1 envelope V3 loop was deep-sequenced from pre-transplant PBMC DNA. Computational algorithms (e.g., geno2pheno) were used to predict whether the patient's archived virus was CCR5-tropic or CXCR4-tropic [55].
- Ex Vivo Viral Challenge: Post-transplant CD4+ T cells from the patient were isolated and challenged in vitro with:
- CCR5-tropic viruses (e.g., Ba-L, ZM247)
- CXCR4-tropic virus (e.g., NL4-3) Productive infection was measured via intracellular p24 staining and infectivity of culture supernatants on indicator cell lines over 7 days [55]. Cells from the London Patient were resistant to CCR5-tropic virus but susceptible to CXCR4-tropic virus, confirming a lack of CCR5 expression.
Immunological and Serological Assays
- HIV-Specific T-Cell Responses: Intracellular cytokine staining (ICS) was performed after stimulation with HIV-1 Gag peptides to measure HIV-1-specific CD4+ and CD8+ T-cell responses. A loss of these responses post-transplant indicates a lack of antigenic stimulation [55] [58].
- Humoral Response: Levels and avidity of HIV-1-specific antibodies (e.g., against Env) were measured. A steady decline in antibody titers and avidity over time is indicative of the absence of viral replication [55] [58].
Diagram 1: Experimental Workflow for Validating HIV Cure Post-Transplant
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents and Assays
| Reagent / Assay | Function / Application | Example Use in Case Studies |
|---|---|---|
| Droplet Digital PCR (ddPCR) | Absolute quantification of HIV-1 DNA (LTR, Gag) with high precision and sensitivity to measure reservoir size. | Quantified total HIV-1 DNA in CD4+ T cells of the London Patient [58]. |
| Quantitative Viral Outgrowth Assay (QVOA) | Gold-standard functional assay to quantify the replication-competent latent reservoir in resting CD4+ T cells. | Used on 24 million resting CD4+ T cells from the London Patient; no virus detected [55]. |
| Single Genome Sequencing (SGS) | Amplifies and sequences individual HIV-1 envelope genes to determine viral tropism without recombination. | Performed on pre-transplant PBMC DNA to confirm CCR5-tropism of the archived virus [55]. |
| Flow Cytometry with p24 ICS | Detects intracellular HIV-1 p24 antigen to confirm active infection in cells after in vitro stimulation or challenge. | Used in ex vivo viral challenge assays to confirm lack of CCR5-tropic infection post-transplant [55]. |
| Chimerism Analysis | Measures the proportion of donor-derived cells (e.g., in whole leukocytes or T-cell subsets) in the recipient. | Confirmed >99% donor chimerism in the London Patient's T-cells [58]. |
| Alemtuzumab (Anti-CD52) | In vivo T-cell depletion antibody used in conditioning regimens to prevent graft rejection and GvHD. | Part of the reduced-intensity conditioning for the London Patient [55]. |
Mechanistic Insights and Signaling Pathways
The cure in these patients is a multi-factorial process involving the replacement of susceptible host cells with HIV-resistant ones and potential immune-mediated clearance of the reservoir.
Diagram 2: Post-Transplant Mechanisms Leading to HIV Cure
The primary mechanism of cure is the replacement of the susceptible host immune system with donor-derived CCR5-negative cells, creating an environment where the patient's predominant R5-tropic virus cannot establish new infections [55] [56]. The conditioning regimen (chemotherapy/radiotherapy) ablates a significant portion of the existing reservoir [57]. Furthermore, a graft-versus-reservoir effect, analogous to graft-versus-leukemia, is hypothesized to occur, where the new donor immune system recognizes and clears remaining HIV-infected host cells [57]. Recent research on a subsequent patient (the "Second Berlin Patient") highlights a potential role for unusual natural killer (NK) cell responses in directing antibodies to clear the residual reservoir via antibody-dependent cellular cytotoxicity (ADCC) [59].
Beyond the Cases: Implications for Future Research and Therapy
The lessons from these patients have directly informed the development of several promising therapeutic avenues aimed at achieving a more widely applicable HIV cure or long-term remission.
- Gene Editing Strategies: Technologies like CRISPR/Cas9, ZFNs, and TALENs are being explored to disrupt the CCR5 gene in a patient's own cells, mimicking the CCR5Δ32 phenotype [3]. Clinical trials (e.g., NCT03164135) have demonstrated the feasibility and safety of CRISPR/Cas9-mediated CCR5 editing in hematopoietic stem cells [3]. To counter the potential for viral escape, multiplex gene editing strategies that simultaneously target CCR5, CXCR4, and the HIV-1 proviral LTR are under investigation [3].
- Synergistic Immunotherapy: Combining gene editing with immune-based approaches is a frontier in cure research. This includes engineering CAR-T cells that target HIV-infected cells, and using immune checkpoint inhibitors to reverse T-cell exhaustion and enhance the clearance of the latent reservoir [3].
- Expanding the Donor Pool: The case of the "Second Berlin Patient," who was cured after a transplant from a heterozygous (CCR5Δ32/wt) donor, suggests that complete CCR5 ablation may not be strictly necessary [57] [59]. This significantly expands the potential donor pool for such procedures and lowers the barrier for gene therapy strategies, which may only need to achieve a certain threshold of CCR5 disruption to be effective.
The cases of the Berlin and London Patients stand as transformative proof-of-concept that HIV-1 can be cured. They unequivocally validated CCR5 as a critical therapeutic target and demonstrated that replacing a patient's immune system with a CCR5-deficient one can lead to sustained viral remission. The extensive experimental protocols developed to verify their cure now serve as a rigorous benchmark for the field. While allo-HSCT itself is not a scalable cure, the insights gleaned have catalyzed the development of sophisticated and potentially more accessible next-generation therapies, including gene editing and combinatorial immunotherapies, bringing the goal of a widely available HIV cure closer to reality.
The pursuit of a sterilizing or functional cure for Human Immunodeficiency Virus (HIV) represents a paramount challenge in modern medical science. While highly active antiretroviral therapy (ART) can effectively suppress viral replication, it fails to eradicate latent viral reservoirs and necessitates lifelong adherence, leading to cumulative drug toxicity and the potential emergence of resistant strains [21] [3]. The discovery of C-C chemokine receptor 5 (CCR5) as a crucial co-receptor for HIV entry marked a turning point, a fact powerfully illustrated by the "Berlin" and "London" patients who achieved sustained viral remission after receiving hematopoietic stem cell transplants from donors with a homozygous CCR5-Δ32 mutation [21] [3]. This natural resistance to R5-tropic HIV strains provided a compelling therapeutic rationale for disrupting CCR5 via gene editing. However, the high mutability of HIV allows for viral escape, notably through a tropism switch to employ the CXCR4 co-receptor [21]. Furthermore, the integrated provirus within the host genome can exploit the viral Long Terminal Repeat (LTR) region to reactivate from latency [21]. Consequently, single-target strategies are insufficient. This whitepaper elucidates the advanced strategy of multiplexed gene editing—simultaneously targeting CCR5, CXCR4, and HIV LTR—to establish a comprehensive viral blockade, thereby countering tropism switching, preventing latent reactivation, and offering a pathway to a definitive HIV cure [21].
Biological Foundations and Therapeutic Targets
CCR5: The Primary HIV Co-receptor
For the predominant R5-tropic HIV-1 strains, CCR5 is an essential co-receptor for viral entry into CD4+ T cells and macrophages [21]. Its expression on the cell surface directly determines cellular susceptibility to infection. The CCR5-Δ32 mutation, a natural 32-base-pair deletion that results in a non-functional receptor, confers high resistance to HIV-1 infection in homozygous individuals, validating CCR5 as a prime therapeutic target [21] [3].
CXCR4: The Alternate Co-receptor and Vector for Viral Escape
Following successful CCR5 disruption, selective pressure can facilitate the outgrowth of existing or emergent X4-tropic HIV strains that utilize the CXCR4 co-receptor for cell entry [21]. This tropism switching allows the virus to circumvent CCR5-targeted therapies and sustain infection. Simultaneous knockout of CXCR4 is therefore critical to prevent this escape route and establish a robust barrier against both major viral tropisms [21].
HIV LTR: The Master Regulator of Viral Latency and Reactivation
The integrated HIV provirus remains the primary obstacle to a cure. The viral Long Terminal Repeat (LTR) regions contain powerful promoter and enhancer elements that govern viral gene expression [21]. Even in cells lacking CCR5 or CXCR4, the latent provirus can be reactivated via the LTR, leading to viral rebound upon therapy cessation. Key functions of the LTR include:
- Transcriptional Activation: Serving as the primary site for initiation of viral RNA synthesis [21].
- Gag Expression and Viral Assembly: LTR-driven transcription is essential for the production of structural proteins like Gag and the assembly of new viral particles [21].
- Reverse Transcription and Integration: The LTR facilitates reverse transcription and is the site for integrase-mediated insertion into the host genome [21]. Gene editing aimed at disrupting the LTR can permanently silence the provirus, preventing reactivation from the latent reservoir and achieving a functional cure.
Gene Editing Technologies for HIV Therapy
The development of precise molecular tools has enabled the targeted editing of host and viral genomes. The comparative characteristics of major platforms are summarized in Table 1.
Table 1: Comparative Characteristics of Major Gene Editing Technologies for HIV Therapy
| Technology | Mechanism of Action | Advantages | Limitations and Challenges | Representative Advances in HIV |
|---|---|---|---|---|
| Zinc Finger Nucleases (ZFNs) | Custom-designed zinc finger proteins fuse with FokI nuclease to induce DNA cleavage. | Early clinical data; first CCR5-editing technology in clinical trials. | Complex design; higher off-target risk; potential immunogenicity. | SB-728-T clinical trial showed safety and virological/immunological benefits [3]. |
| TALENs | Transcription activator-like effector proteins fuse with FokI nuclease for DNA cleavage. | Modular design offers better specificity than ZFNs. | Technically demanding; large size hinders viral delivery. | Automated, clinical-scale production of TALEN-edited CD4+ T cells developed [21] [3]. |
| CRISPR/Cas9 | A single guide RNA (sgRNA) directs Cas9 nuclease for site-specific DNA double-strand breaks. | Easy design; high efficiency; excellent multiplex capability. | Off-target effects; PAM sequence dependency; potential immune response. | High efficiency in models; early-phase clinical trials (e.g., NCT03164135) demonstrate feasibility [21] [3]. |
| Base Editors (BE) | Cas9 nickase fused to a deaminase enables precise single-nucleotide changes without double-strand breaks. | Avoids double-strand break risks (indels, translocations). | Off-target DNA/RNA editing; limited editing window. | PD-1 gene edited using CE-8e-SpRY mRNA delivered via lentiviral-like particles [21] [3]. |
Quantitative Analysis of Editing Efficiency and Safety
A critical step in therapeutic development is the empirical evaluation of editing platforms. Performance metrics for key systems are detailed in Table 2.
Table 2: Quantitative On-Target Efficiency and Off-Target Profiles of Gene Editing Platforms
| Editing Platform | Target Locus | Reported On-Target Efficiency (%) | Off-Target Risk Profile | Key Mitigation Strategies |
|---|---|---|---|---|
| CRISPR/Cas9 | CCR5 | 60-85 [21] | Medium to High | High-fidelity Cas9 variants; optimized sgRNA design [21]. |
| CRISPR/Cas9 | CXCR4 | 70-90 [21] | Medium to High | Truncated sgRNAs; dual nickase strategy [21]. |
| TALENs | CCR5 | 40-75 [21] | Low to Medium | Protein-driven specificity reduces off-targets [21]. |
| Base Editors | PD-1 (as a model) | >80 [21] | Low (for indels) | The absence of double-strand breaks minimizes chromosomal abnormalities [21]. |
Experimental Workflow for Multiplexed Gene Editing
The following diagram outlines a standardized experimental protocol for implementing and validating a multiplexed editing strategy in a research setting.
Diagram 1: Workflow for multiplexed gene editing experiments.
Detailed Experimental Protocols
5.1.1 sgRNA Design and Vector Construction for Multiplexed Editing
- Design Principles: For CRISPR/Cas9, design sgRNAs (typically 20 nt) with high on-target scores and minimal off-target potential for human CCR5 (chr3:46373440..46379238), CXCR4 (chr2:136114293..136118159), and conserved regions of the HIV-1 LTR. Computational tools like CRISPRscan and BLAST against the human genome are essential [21].
- Multiplex Cloning Strategy: Utilize a Csy4 endonuclease-based system for processing multiple gRNAs from a single transcript [60]. Clone gRNA sequences under the control of the RNA polymerase III promoter (e.g., U6 or SNR52p) with a SUP4t terminator into a high-copy 2μ plasmid. A Csy4 gene, driven by a constitutive promoter like PGK1p, should be co-expressed to release individual gRNAs [60].
- Delivery Vector Assembly: For in vivo applications, package the multiplex gRNA construct and the Cas9 gene (e.g., SpCas9 or high-fidelity eSpCas9) into a lentiviral vector. Employ a "Gag-only" packaging strategy with lentiviral-like particles (LVLPs) to enhance safety by minimizing the generation of replication-competent virus [21] [3].
5.1.2 Cell Transduction and Analysis of Editing Efficiency
- Target Cells: Primary human CD4+ T cells or hematopoietic stem/progenitor cells (HSPCs) are isolated from donors. For latency models, use cell lines with integrated HIV provirus (e.g., J-Lat cells).
- Transduction: Activate T cells with CD3/CD28 beads and transduce with lentiviral vectors at a pre-optimized Multiplicity of Infection (MOI). For HSPCs, use spinoculation to enhance transduction efficiency.
- Efficiency Measurement: 72-96 hours post-transduction, harvest cells. Use Tracking of Indels by Decomposition (TIDE) or next-generation sequencing (NGS) of PCR-amplified target loci to quantify insertion/deletion (indel) frequencies for CCR5 and CXCR4. For LTR disruption, employ a combination of NGS and a reduction in LTR-driven reporter gene expression (e.g., GFP) upon cell activation.
5.1.3 Functional Validation of HIV Inhibition
- Viral Challenge Assay: Challenge edited primary CD4+ T cells with R5-tropic (e.g., Bal.), X4-tropic (e.g., NL4-3), and dual-tropic HIV-1 strains. Monitor viral replication over 10-14 days by measuring p24 antigen levels in the supernatant via ELISA.
- Latency Reactivation Assay: Treat edited latent cell models (e.g., J-Lat) with latency-reversing agents (LRAs) like PMA/ionomycin or TNF-α. Quantify the percentage of GFP+ cells via flow cytometry or measure viral RNA output to assess the success of LTR disruption in preventing reactivation.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions for Multiplexed Gene Editing Studies
| Reagent / Tool | Function / Application | Example / Specification |
|---|---|---|
| CRISPR/Cas9 System | Core editing machinery for inducing targeted DNA breaks. | High-fidelity SpCas9 (eSpCas9(1.1)); Cas12a (Cpf1) for multiplexing with crRNA arrays [21] [60]. |
| Multiplex gRNA Vector | Simultaneously expresses multiple guide RNAs from a single construct. | Plasmid with SNR52p promoter and Csy4 ribozyme sites for processing individual gRNAs [60]. |
| Lentiviral-Like Particles (LVLPs) | Safe and efficient delivery of editor mRNA/gRNA in vivo. | "Gag-only" packaging system to deliver CE-8e-SpRY base editor mRNA [21] [3]. |
| Humanized Mouse Models | In vivo testing of edited cell efficacy and HIV challenge. | NSG mice engrafted with human immune systems (e.g., CD34+ HSPCs) [3]. |
| Next-Gen Sequencing (NGS) | Comprehensive analysis of on-target efficiency and genome-wide off-target effects. | Targeted amplicon sequencing for on-target; GUIDE-seq or CIRCLE-seq for unbiased off-target discovery. |
Integrated Strategy: Synergy with Immunotherapy
Multiplexed gene editing alone may not fully clear existing reservoirs. A synergistic combination with immunotherapy is crucial. The engineered cells can be empowered to not only resist new infection but also actively eliminate infected cells, as illustrated below.
Diagram 2: Gene editing and immunotherapy synergy.
This synergy can be operationalized through several advanced modalities:
- Armored CAR-T Cells: Generate HIV-specific CAR-T cells (e.g., targeting the HIV envelope) and simultaneously edit their CCR5 and CXCR4 genes to protect them from infection during their anti-viral activity. These cells can be further engineered to secrete PD-1-blocking single-chain variable fragments (scFvs), which can reverse T-cell exhaustion in the local microenvironment and enhance the killing of HIV-infected cells [21] [3].
- Combination with Immune Checkpoint Inhibitors: Systemic or targeted administration of antibodies against PD-1 or other checkpoint molecules can reinvigorate exhausted HIV-specific CD8+ T cells. When combined with multiplex gene editing in hematopoietic stem cells, this approach can generate a protected and rejuvenated immune system capable of controlling or eliminating the virus [21].
- Broadly Neutralizing Antibodies (bNAbs): Infusions of bNAbs can clear circulating virus and opsonize infected cells. A new wave of protected, edited CD4+ T cells emerging from transplanted HSPCs can help rebuild a functional immune system that is resistant to re-infection, creating a favorable environment for long-term remission [21].
Multiplexed gene editing, which simultaneously targets the host factors CCR5 and CXCR4 and the viral HIV LTR, represents a paradigm shift in the pursuit of an HIV cure. This strategy constructs a multi-layered defense that effectively blocks viral entry, preempts viral escape via tropism switching, and permanently silences the integrated provirus. When synergistically combined with innovative immunotherapies such as armored CAR-T cells and immune checkpoint modulation, this approach promises to overcome the limitations of current ART by enabling the generation of a virus-resistant immune system capable of durable viral control. While significant challenges regarding long-term safety, delivery efficiency, and global accessibility remain, the integrated framework of multiplexed gene editing and immunotherapy provides a robust and promising foundation for the next generation of HIV treatment paradigms [21] [3].
The convergence of cell engineering, gene editing, and immunotherapy represents a paradigm shift in the treatment of persistent viral infections and cancer. This whitepaper examines the synergistic potential of combining CCR5-edited chimeric antigen receptor (CAR)-T cells with immune checkpoint inhibitors, a strategy grounded in foundational research on the CCR5-Δ32 mutation's role in conferring natural resistance to HIV-1 infection. We provide a technical overview of the underlying mechanisms, detail experimental protocols for developing these advanced cellular products, and summarize quantitative data comparing gene-editing platforms. Furthermore, we visualize key signaling pathways and experimental workflows and catalog essential research reagents to facilitate the translation of this integrated approach from concept to clinical application.
The chemokine receptor CCR5, a G protein-coupled receptor (GPCR) expressed on the surface of immune cells such as T lymphocytes, macrophages, and dendritic cells, serves as the predominant co-receptor for human immunodeficiency virus (HIV) entry during initial transmission and early stages of infection [18] [11]. The discovery that a homozygous 32-base pair deletion (CCR5-Δ32) in the CCR5 gene prevents cell surface expression and confers profound resistance to CCR5-tropic HIV strains provided a genetic blueprint for a curative strategy [18] [11]. This natural experiment was validated by the cases of the "Berlin" and "London" patients, who achieved sustained viral remission after allogeneic hematopoietic stem cell transplantation from CCR5-Δ32/Δ32 donors, proving that CCR5 disruption can functionally cure HIV [3].
Beyond its role in HIV entry, CCR5 mediates the recruitment of immune cells to sites of inflammation and is implicated in cancer metastasis and the regulation of immunosuppressive cell populations within the tumor microenvironment (TME) [61] [11]. Consequently, targeting CCR5 has therapeutic relevance that extends from infectious diseases to oncology. The integration of CCR5 disruption into CAR-T cell engineering creates a multifaceted therapeutic platform. These engineered cells are designed not only to target specific antigens via their CAR but also to resist HIV-mediated destruction and potentially enhance their trafficking and function within immunosuppressive microenvironments, particularly when combined with checkpoint inhibitors that reverse T-cell exhaustion [3] [62].
Scientific Rationale for a Synergistic Approach
The Dual Role of CCR5 in HIV Infection and Cancer Immunobiology
CCR5's role as a HIV co-receptor makes it a critical target for protecting immune cells from infection. HIV entry into CD4+ T cells requires sequential binding of the viral envelope glycoprotein gp120 to the CD4 receptor followed by interaction with CCR5 (or CXCR4), which triggers fusion and viral entry [11] [1]. Disrupting this interaction is a validated therapeutic strategy.
Simultaneously, in oncology, the CCR5/CCL5 axis is hijacked by tumors to promote metastasis and maintain an immunosuppressive TME. CCR5 activation on Tregs augments their differentiation and migration to tumor sites, while its expression on cancer cells activates calcium signaling and downstream pathways like PI3K/AKT, inducing cell survival, resistance to DNA-damaging agents, and a "stemness" phenotype [61]. Table 1 summarizes the multifaceted roles of CCR5 across disease contexts.
Table 1: Multifaceted Roles of CCR5 in Disease and Therapy
| Context | Role of CCR5 | Therapeutic Implication |
|---|---|---|
| HIV Infection | Primary co-receptor for viral entry into CD4+ T cells [18] [11] | Disruption confers resistance to R5-tropic HIV infection. |
| Cancer Metastasis | Induces cancer cell homing to metastatic sites; augments invasiveness [61] | CCR5 inhibition (e.g., with maraviroc) can restrain cancer metastasis. |
| Immune Suppression | Recruits Tregs and MDSCs to the TME; expressed preferentially on Tregs [61] [62] | Blocking CCR5 may reduce immunosuppressive cell populations in TME. |
| Therapeutic Resistance | Enhances DNA repair; confers resistance to DNA-damaging agents [61] | CCR5 inhibitors may sensitize tumors to conventional therapies. |
Mechanisms of Synergy: CCR5 Editing, CAR-T Cells, and Checkpoint Inhibition
The synergy of this combination therapy arises from targeting complementary immune evasion pathways. First, CCR5 editing intrinsically protects CAR-T cells from HIV infection, which is crucial for maintaining a durable therapeutic cell population in HIV/AIDS patients, including those with HIV-associated malignancies [3] [63]. Furthermore, CCR5 disruption may alter the functional properties of T cells, potentially reducing exhaustion phenotypes and improving persistence.
Second, combining this approach with checkpoint inhibitors addresses the profound T-cell exhaustion characteristic of both chronic HIV infection and cancer. The persistent antigen exposure in these conditions leads to upregulation of inhibitory receptors like PD-1, which suppresses T-cell effector functions [62]. Checkpoint inhibitors (e.g., anti-PD-1 antibodies) block these interactions, "releasing the brakes" on T cells and restoring their cytotoxic capacity. When applied to CCR5-edited CAR-T cells, this combination can theoretically reverse the exhaustion of both the engineered and endogenous T cells, leading to a more robust and sustained anti-tumor or anti-viral response [3] [62].
Diagram 1: Logical workflow illustrating the problem of immune dysfunction in chronic HIV/cancer and the synergistic mechanism of the combined immunotherapy. CCR5 editing addresses susceptibility to HIV and the TME, while checkpoint inhibition counters T-cell exhaustion, leading to a potent, durable immune response.
Experimental Protocols and Methodologies
Protocol for CCR5 Gene Editing in Primary Human T Cells
This protocol outlines the process for using CRISPR/Cas9 to disrupt the CCR5 gene in primary human T cells, a critical first step in generating HIV-resistant CAR-T cells [3] [63].
Materials:
- Source Cells: Isolated human peripheral blood mononuclear cells (PBMCs) or purified CD4+/CD8+ T cells from leukapheresis product.
- Activation: Anti-CD3/CD28 activation beads.
- Culture Media: X-VIVO 15 or RPMI-1640 supplemented with IL-2 (100-300 IU/mL) and 10% FBS.
- Gene Editing Components:
- RNP Complex: Recombinant S. pyogenes Cas9 protein and synthetic sgRNA targeting CCR5 (e.g., 5'-GAG AAG TGT CAG TTC ATA CCC-3').
- Delivery Method: Electroporation system (e.g., Lonza 4D-Nucleofector).
- Analysis: Flow cytometry antibodies for CCR5 (CD195), CD3, CD4, CD8; T7E1 assay or next-generation sequencing for indel analysis.
Procedure:
- T Cell Isolation and Activation: Isolate T cells from PBMCs using a negative selection kit. Activate the T cells using anti-CD3/CD28 beads at a 1:1 bead-to-cell ratio in complete media supplemented with IL-2 for 24-48 hours.
- Ribonucleoprotein (RNP) Complex Formation: Complex the Cas9 protein (30-60 pmol) with the sgRNA (60-120 pmol) targeting CCR5 in a sterile tube. Incubate at room temperature for 10-20 minutes to form the RNP complex.
- Electroporation: Harvest activated T cells, wash with PBS, and resuspend in the appropriate electroporation buffer. Combine the cell suspension with the pre-formed RNP complex and transfer to an electroporation cuvette. Electroporate using a pre-optimized program (e.g., EO-115 on the 4D-Nucleofector).
- Recovery and Expansion: Immediately after electroporation, transfer cells to pre-warmed complete media with IL-2. Culture the cells at 37°C, 5% CO2. Remove activation beads after 7-10 days and expand cells as needed.
- Assessment of Editing Efficiency: 3-5 days post-electroporation, analyze a sample of cells by flow cytometry to assess the loss of CCR5 surface expression. Genomically, extract DNA and use the T7E1 assay or NGS of the targeted CCR5 locus to quantify the frequency of insertions/deletions (indels). Expect editing efficiencies of 50-80% with optimized protocols.
Protocol for Combining CCR5-Edited CAR-T Cells with Checkpoint Inhibitors In Vivo
This protocol describes the key steps for evaluating the efficacy of the combination therapy in an immunodeficient mouse model of human cancer [3] [62].
Materials:
- Animals: NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (NSG) mice, 6-8 weeks old.
- Cancer Cell Line: A relevant human cancer cell line (e.g., CCR5+ solid tumor line).
- Therapeutic Cells: CCR5-edited CAR-T cells (vs. non-edited CAR-T controls).
- Checkpoint Inhibitor: Clinical-grade anti-human PD-1 monoclonal antibody (e.g., Nivolumab biosimilar).
- In Vivo Imaging System (IVIS) for monitoring tumor growth if using luciferase-expressing cells.
Procedure:
- Tumor Engraftment: Subcutaneously inject the human cancer cells (e.g., 5x10^6 cells in Matrigel) into the flanks of NSG mice. Allow tumors to establish to a palpable size (~50-100 mm³).
- Cell Therapy Administration: Randomize mice into four treatment groups: (i) Untreated control, (ii) Anti-PD-1 alone, (iii) CCR5-edited CAR-T cells alone, (iv) CCR5-edited CAR-T cells + anti-PD-1. Intravenously inject a therapeutic dose of CAR-T cells (e.g., 5-10x10^6 cells per mouse).
- Checkpoint Inhibitor Dosing: Administer the anti-PD-1 antibody (e.g., 200 µg per dose) via intraperitoneal injection. Begin treatment shortly after CAR-T cell infusion (e.g., day 1) and continue for multiple doses (e.g., twice weekly for 2-3 weeks).
- Monitoring and Endpoint Analysis:
- Tumor Volume: Measure tumor dimensions 2-3 times weekly with calipers. Calculate volume as (length x width²)/2.
- Body Weight: Monitor as an indicator of overall health and toxicity.
- In Vivo Bioluminescence: If using luciferase-expressing CAR-T cells, inject luciferin substrate and image weekly to track CAR-T cell persistence and trafficking.
- Endpoint Immunophenotyping: At the study endpoint, harvest tumors and spleen. Generate single-cell suspensions and analyze by flow cytometry for tumor-infiltrating lymphocytes (TILs): quantify human CD3+ CAR+ T cells, effector memory subsets, and exhaustion markers (PD-1, LAG-3, TIM-3).
Quantitative Data and Technology Comparison
The selection of an appropriate gene-editing platform is critical for the successful development of CCR5-edited CAR-T cells. Table 2 provides a comparative analysis of the primary technologies in use.
Table 2: Comparison of Gene-Editing Platforms for CCR5 Disruption
| Technology | Mechanism of Action | Editing Efficiency | Key Advantages | Key Limitations & Challenges |
|---|---|---|---|---|
| ZFNs | Custom zinc-finger proteins fused to FokI nuclease dimer induce DNA cleavage [3]. | Moderate | First platform in clinical trials (SB-728-T); established clinical safety/efficacy data [3]. | Complex design/construction; higher risk of off-target effects; potential immunogenicity [3]. |
| TALENs | Transcription activator-like effector proteins fused to FokI nuclease dimer induce DNA cleavage [3]. | Moderate | More modular than ZFNs; improved specificity and reduced off-target risk vs. ZFNs [3]. | Technically demanding construction; large size hinders viral vector delivery [3]. |
| CRISPR/Cas9 | sgRNA directs Cas9 nuclease to genomic locus for a double-strand break [3]. | High (Typically >60% in T cells) | Easy design; high efficiency; allows multiplexed editing of several genes [3] [64]. | Off-target effects are main safety concern; PAM sequence dependency; long-term immune response to Cas9 [3]. |
| Base Editors | Cas9 nickase fused to deaminase enables precise single-nucleotide conversion without double-strand breaks [3]. | High (Dependent on target base) | Avoids risks of double-strand breaks (indels, chromosomal translocations); high precision [3]. | Potential for off-target DNA/RNA editing; limited editing window constrains targetable sites [3]. |
Recent clinical trials underscore the translational potential of these platforms. The clinical trial NCT03164135 assessed CRISPR/Cas9-mediated CCR5 editing in hematopoietic stem cells for patients with HIV and acute lymphoblastic leukemia, demonstrating feasibility and a acceptable safety profile [3]. Furthermore, a study utilizing ZFN-modified CCR5-autologous T cells (SB-728-T) demonstrated that the approach could lead to significant reductions in viral load and immunological benefits [3].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions for Developing Combination Immunotherapy
| Research Reagent / Tool | Function & Application | Example Products / Assays |
|---|---|---|
| CCR5 Gene Editing Tools | Precise disruption of the CCR5 gene in T cells to confer HIV resistance. | CRISPR/Cas9 RNP (e.g., Alt-R S.p. Cas9), ZFNs (e.g., SB-728-T), TALENs [3]. |
| Viral Vector Systems | Stable delivery of CAR transgenes and editing machinery into T cells. | Lentiviral vectors, Retroviral vectors (e.g., for CAR delivery) [64]. |
| CAR-T Cell Culture Supplements | Support T-cell activation, survival, and expansion during ex vivo manufacturing. | Anti-CD3/CD28 activation beads, recombinant human IL-2, IL-7, IL-15 [65] [63]. |
| Checkpoint Inhibitors | Block inhibitory receptors on T cells to reverse exhaustion and enhance function. | Anti-PD-1 (Nivolumab, Pembrolizumab), Anti-PD-L1 (Atezolizumab) [62]. |
| In Vivo Model Systems | Pre-clinical evaluation of therapy efficacy, persistence, and safety. | NSG (NOD-scid-gamma) mice, patient-derived xenograft (PDX) models [61]. |
| Multiparameter Flow Cytometry Panels | Analyze cell surface markers, CAR expression, intracellular cytokines, and exhaustion profiles. | Antibodies against CD3, CD4, CD8, CCR5, PD-1, LAG-3, TIM-3, IFN-γ, TNF-α [63] [62]. |
Signaling Pathways and Molecular Interactions
The synergistic effect of this combination therapy operates through interconnected signaling networks. The diagram below maps the key pathways involved in CCR5 function, CAR signaling, and PD-1 mediated inhibition.
Diagram 2: Key signaling pathways in synergistic immunotherapy. CCR5 activation triggers chemotaxis and cell migration pathways, which are hijacked by HIV for entry. The CAR signal drives T-cell activation and cytotoxicity. The PD-1 pathway inhibits T-cell function, an effect reversed by checkpoint inhibitors. Cross-talk, such as CCR5-mediated PI3K/AKT signaling, can augment CAR-T cell survival.
The C-C chemokine receptor 5 (CCR5) serves as the primary coreceptor for human immunodeficiency virus (HIV) entry into CD4+ T lymphocytes, making it a critical target for therapeutic intervention [11]. The discovery that individuals carrying a homozygous 32-base pair deletion in the CCR5 gene (CCR5Δ32) demonstrate near-complete resistance to HIV infection established the foundational premise for targeting this receptor [34] [12]. This scientific rationale has since driven the development of various CCR5-targeted therapies, necessitating robust and predictive in vitro and in vivo models to assess their efficacy from the laboratory bench to clinical application. This whitepaper provides a comprehensive technical guide to the experimental models employed in this field, detailing their applications, methodologies, and integration into the therapeutic development pipeline.
In Vivo Model Systems
Humanized Mouse Models
Humanized mice, created by engrafting human hematopoietic tissues or cells into immunodeficient mice, provide a critical in vivo platform for studying HIV pathogenesis and evaluating therapeutic interventions.
- Model Generation: The most common approach involves transplanting human CD34+ hematopoietic stem cells (HSCs) into severely immunodeficient mice (e.g., NSG or NOG strains). Over approximately 12 weeks, these cells reconstitute a human immune system, including T cells, B cells, and macrophages, within the murine host [66].
- Application in CCR5 Research: These models are indispensable for testing the efficacy of CCR5 gene editing strategies. For instance, Khamaikawin et al. (2018) used humanized mice to model the bone marrow transplantation strategy of the "Berlin patient" [66]. After establishing HIV infection, mice underwent bone marrow ablation and secondary transplantation with a mixture of HIV-resistant and susceptible HSCs. The study demonstrated that HIV-resistant CD4+ cells derived from the edited HSCs selectively expanded in the presence of the virus, recapitulating the human clinical outcome [66].
- Recent Advancements: A landmark 2025 study published in Nature Communications demonstrated high-frequency (>90%) CCR5 editing in human mobilized CD34+ HSCs using CRISPR/Cas9. These edited cells were transplanted into xenograft mice, which subsequently developed a human immune system largely refractory to HIV infection. The study provided crucial pre-clinical proof-of-concept for an autologous, CCR5-knockout hematopoietic stem cell transplant (HSCT) as a curative strategy [22].
The diagram below illustrates the typical workflow for evaluating CCR5-edited therapies in humanized mouse models.
Threshold of Protection and Titration Studies
A key finding from humanized mouse studies is the existence of a threshold effect for CCR5 disruption. The 2025 Nature Communications study systematically titrated the percentage of CCR5-edited HSPCs in the transplant infusion and challenged the mice with HIV [22]. The results demonstrated that:
- High Protection: ≥90% CCR5 editing conferred robust protection against HIV infection.
- Decreasing Benefit: Editing frequencies below 90% provided diminishing protective benefit.
- Negligible Protection: The protective effect became negligible when editing frequencies fell between 54% and 26% [22].
This threshold effect explains the failure of allogeneic HSCT with CCR5WT/Δ32 heterozygous cells (which provide only 50% theoretical disruption) to prevent HIV rebound and underscores the necessity of achieving high editing efficiency for a functional cure [22].
In Vitro Model Systems
In vitro models provide a controlled environment for initial efficacy and safety testing.
Primary Cell Cultures
- Target Cells: Primary human CD4+ T lymphocytes or CD34+ Hematopoietic Stem and Progenitor Cells (HSPCs) are isolated from peripheral blood or mobilized donors [22].
- Editing Protocol: Cells are electroporated with CRISPR/Cas9 ribonucleoproteins (RNPs) complexed with guide RNAs (gRNAs) targeting CCR5.
- Efficacy Assessment: Editing efficiency is quantified via DNA sequencing (indel frequency) and flow cytometry for CCR5 surface expression. The functionality of edited cells is tested by challenging them with CCR5-tropic HIV (e.g., HIVJRCSF) and monitoring infection through p24 ELISA or intracellular viral RNA staining [22].
Guide RNA Selection and Validation
The discovery of optimal gRNAs is a critical, multi-step process as detailed in recent research [22]:
- In Silico Prediction: Software identifies ~123 gRNAs targeting exon 3 of the CCR5 open reading frame.
- Off-Target Filtering: gRNAs with potential binding sites elsewhere in the human genome (e.g., homologous CCR2 gene) are excluded.
- In Vitro Screening: The top candidates are tested in primary human HSPCs. In one study, four gRNAs (TB7, TB8, TB48, TB50) achieved >30% editing frequency with minimal off-target effects.
- Potency Validation: The most potent gRNAs (TB48, TB50) and a dual-guide approach (TB48+TB50) demonstrated superior reduction of CCR5+ CD4+ T cells and conferred the highest level of HIV resistance in vitro [22].
Clinical Trials: From Models to Human Application
Clinical trials represent the ultimate validation of preclinical findings. The following table summarizes key quantitative data from clinical and pre-clinical studies involving CCR5 modulation.
Table 1: Quantitative Efficacy Outcomes of CCR5-Targeted Approaches in Models and Clinical Trials
| Model / Trial | CCR5-Targeting Approach | Key Efficacy Metric | Result | Source |
|---|---|---|---|---|
| Humanized Mice | CRISPR/Cas9 (gRNAs TB48/TB50) in HSPCs | CCR5 Editing Frequency | 91-97% | [22] |
| Humanized Mice | Titrated CCR5-edited HSPCs | Threshold for HIV Protection | ≥90% | [22] |
| In Vitro T Cells | CRISPR/Cas9 (gRNAs TB48/TB50) | Reduction in HIV infection (AUC) | Significantly Reduced | [22] |
| Clinical Case (Berlin Patient) | Allogeneic CCR5Δ32/Δ32 HSCT | HIV Remission | >10 years | [66] [3] |
| Clinical Trial | CRISPR/Cas9 in HSPCs (NCT03164135) | Feasibility & Safety | Demonstrated | [3] [21] |
| ZFN Clinical Trial (SB-728-T) | ZFN-edited autologous T-cells | Safety & Immunological Benefit | Acceptable Profile | [3] |
Essential Research Reagents and Protocols
Table 2: The Scientist's Toolkit: Key Reagents for CCR5/HIV Model Research
| Research Reagent | Function and Role in Research | Example Application |
|---|---|---|
| Immunodeficient Mice | Host for engraftment of human cells; lacks murine immune system to prevent rejection. | NSG, NOG strains for generating humanized mouse models [66]. |
| Human CD34+ HSPCs | Primary cells capable of reconstituting a human immune system in vivo. | Sourced from cord blood or mobilized peripheral blood for transplantation [22]. |
| CRISPR/Cas9 System | Gene editing tool for precise knockout of the CCR5 gene. | Electroporation of Cas9 protein complexed with CCR5-specific gRNAs into HSPCs [22]. |
| CCR5-tropic HIV | Viral strain that utilizes the CCR5 coreceptor for entry; models primary infection. | HIVJRCSF for in vitro and in vivo challenge experiments [22]. |
| Anti-CCR5 Antibody (2D7) | Monoclonal antibody used to detect CCR5 cell surface expression via flow cytometry. | Quantifying the efficiency of CCR5 knockout after gene editing [22] [67]. |
Detailed Experimental Protocol: CCR5 Editing in Human HSPCs
The following protocol is adapted from a 2025 pre-clinical study [22]:
- HSPC Isolation and Culture: Isolate CD34+ HSPCs from leukapheresis samples of G-CSF-mobilized healthy adult donors using clinical-grade magnetic bead separation. Culture cells in serum-free medium supplemented with cytokines (SCF, TPO, FLT3-L).
- CRISPR/Cas9 RNP Electroporation: Complex chemically synthesized gRNAs (e.g., TB48 and TB50) with SpCas9 protein to form ribonucleoproteins (RNPs). Electroporate the RNPs into HSPCs using a validated clinical-scale electroporation system.
- Assessment of Editing Efficiency: At 48 hours post-electroporation, harvest cells.
- Genomic DNA Analysis: Extract genomic DNA and amplify the CCR5 target region by PCR. Use deep sequencing to quantify the frequency of indels and large deletions.
- Flow Cytometry: Stain cells with an anti-CCR5 antibody (e.g., 2D7) to measure the reduction of CCR5 surface expression.
- In Vitro Pluripotency Assay: Perform colony-forming unit (CFU) assays to ensure edited HSPCs retain their ability to differentiate into multiple hematopoietic lineages.
- In Vivo Transplantation and Challenge: Transplant edited HSPCs into sublethally irradiated immunodeficient mice. After 12-16 weeks for immune reconstitution, challenge the mice intraperitoneally with a high dose of CCR5-tropic HIV. Monitor viral load in plasma weekly by RT-PCR and track CD4+ T cell counts by flow cytometry.
Integrated Pathways from Research to Clinical Application
The journey from basic research to clinical application involves a multi-stage process, integrating data from various models, as illustrated below.
Challenges and Future Directions
Despite significant progress, several challenges remain in the use of these models. Humanized mice still exhibit limited functionality of some human immune cell populations and may not fully recapitulate HIV latency in humans [66]. Furthermore, the potential for viral tropism switching—where HIV evolves to use the CXCR4 coreceptor after CCR5 is blocked—necessitates the development of multi-target gene editing strategies targeting both CCR5 and CXCR4 [3] [21]. Future research will focus on improving the safety profile of gene editing, enhancing engraftment efficiency of edited cells, and combining gene editing with immunotherapies (e.g., CAR-T cells) to achieve a synergistic and durable functional cure for HIV [3] [21].
Navigating Hurdles: Addressing Tropism Switching, Safety, and Clinical Heterogeneity
Human Immunodeficiency Virus (HIV) cell entry is a complex process mediated by the interaction between the viral envelope glycoprotein and host cell receptors. The primary receptor is CD4, but a second, crucial co-receptor is absolutely required for viral entry. The two major co-receptors are CCR5 and CXCR4, which are G-protein coupled receptors (GPCRs) normally involved in immune cell signaling and trafficking [68]. Viral tropism refers to the specific co-receptor that a particular HIV strain can utilize. R5-tropic viruses use CCR5, X4-tropic viruses use CXCR4, and dual-tropic (R5X4) viruses can use either co-receptor [69] [70]. The significance of this tropism extends to basic virology and clinical practice. R5-tropic viruses are predominantly responsible for establishing primary infection [71]. However, in approximately 50% of individuals infected with HIV-1 subtype B, a tropism switch occurs during the course of infection, whereby viruses emerge that can use CXCR4 [69] [71]. This switch is clinically critical, as it is strongly associated with an accelerated decline in CD4+ T-cell count and more rapid progression to AIDS [71] [72]. Furthermore, it directly impacts the efficacy of antiretroviral therapy, particularly for the class of drugs known as CCR5 inhibitors [69]. This whitepaper explores the mechanisms, prediction, and clinical consequences of coreceptor tropism switching, framed within the pivotal role of CCR5 and the insights gained from CCR5Δ32 mutation research.
The Biological Foundation of CCR5 and CXCR4
Structural and Functional Biology of Coreceptors
CCR5 and CXCR4 are structurally related members of the large superfamily of G-protein coupled receptors (GPCRs), characterized by seven transmembrane domains [68]. Despite a sequence identity of only 33%, their overall molecular architecture is conserved [73]. A key difference lies in their surface electrostatic potentials: the binding pocket of CXCR4 and its entrance are negatively charged, while CCR5 has a negatively charged pocket but a neutral-to-positive entrance [73]. This difference influences interactions with ligands and the viral envelope.
These receptors bind to endogenous chemokines: CCR5 binds to inflammatory CC-chemokines like CCL3, CCL4, and CCL5, while CXCR4's primary ligand is SDF-1/CXCL12 [68] [73]. The CXCL12/CXCR4 axis is crucial for embryogenesis, hematopoiesis, and neurologesis, as demonstrated by the lethal phenotypic consequences of gene deletion in mice [68]. In contrast, humans with a homozygous CCR5Δ32 mutation, which results in a non-functional CCR5 protein, are generally healthy but exhibit a high level of resistance to HIV-1 infection [68]. This natural resistance highlighted CCR5 as a prime target for antiviral therapy.
The Critical Role of the V3 Loop in Determining Tropism
The viral determinant for coreceptor choice maps largely to the third variable loop (V3) of the gp120 envelope glycoprotein [72] [73]. The V3 loop acts as a key that must fit the lock of the coreceptor. Generally, X4-tropic viruses have V3 sequences with a higher positive net charge, often due to the acquisition of basic amino acids (Lysine or Arginine) at specific positions [72] [73]. The "11/25 rule" is a simple genotypic predictor, which states that the presence of a positively charged amino acid at position 11 or 25 of the V3 loop is indicative of CXCR4 usage [72]. However, tropism switching is a complex process that cannot be explained by V3 sequence alone. Recent evidence shows that mutations outside V3, including in the gp41 region of the envelope, can significantly influence coreceptor use and the efficiency of virus entry [69].
Table 1: Key Characteristics of HIV Coreceptors
| Feature | CCR5 | CXCR4 |
|---|---|---|
| Receptor Family | G-protein Coupled Receptor (GPCR) | G-protein Coupled Receptor (GPCR) |
| Primary Expression | Macrophages, Memory T-cells | Naive T-cells, Broadly expressed |
| Primary Viral Tropism | R5 (Primary/Transmitting virus) | X4 (Emerges later in infection) |
| Endogenous Ligands | CCL3 (MIP-1α), CCL4 (MIP-1β), CCL5 (RANTES) | CXCL12 (SDF-1) |
| Impact of Gene Knockout | Viable; resistant to HIV infection | Lethal in utero (defects in vascular development, hematopoiesis) |
| Key Genotypic Predictor | V3 loop sequence (lower net charge) | V3 loop sequence (higher net charge, 11/25 rule) |
Mechanisms and Drivers of Coreceptor Switching
The Evolutionary Pathway from R5 to X4 Tropism
The transition from exclusive CCR5 use to the emergence of CXCR4-using viruses is a multi-stage evolutionary process within the host. It begins with viruses that still prefer CCR5 but have a nascent, low-efficiency ability to use CXCR4 [69]. These intermediate viruses are often categorized as "dual-R" (CCR5-preferring) [69]. As evolution continues, variants with more robust CXCR4 use ("dual-X") may emerge and eventually become the dominant population. Studies of envelope evolution show that in patients who never undergo a coreceptor switch, entry efficiency via CCR5 continues to improve throughout the infection. In contrast, in patients where a switch is imminent, CCR5 use begins to decline as the virus evolves towards CXCR4 utilization [69].
The Role of Host Immunity and Immune Activation
Viral genetic changes are necessary but not sufficient for a tropism switch; the host's immune environment is a critical facilitator. Recent longitudinal studies have demonstrated that host immune activation is a powerful predictor of a subsequent switch from R5- to X4-tropism [71]. Specifically, elevated levels of activated CD4+ T-cells (measured by %HLA-DR+ CD4+ T-cells and %CD38+HLA-DR+ CD4+ T-cells) measured during early chronic infection independently predict the future emergence of X4-tropic virus [71]. This suggests that an activated immune environment, potentially rich in CXCR4-expressing target cells, creates a fertile ground for X4 variants to emerge and thrive.
The following diagram illustrates the typical progression of HIV infection, highlighting the key virological and immunological events associated with coreceptor switching.
Methodologies for Tropism Determination
Accurate determination of viral tropism is essential for clinical management, especially when considering CCR5 inhibitor therapy. The two main classes of tests are phenotypic and genotypic assays.
Phenotypic Assays
Phenotypic tests directly measure the virus's ability to use CCR5 or CXCR4 to enter cells in a laboratory setting. The Trofile assay (Monogram Biosciences) is a prime example and has been considered a gold standard [69] [72]. These assays are highly accurate but can be time-consuming, expensive, and technically complex, limiting their widespread use in routine clinical practice [72].
Genotypic Prediction and Computational Methods
Genotypic tests infer tropism by analyzing the nucleotide sequence of the V3 loop of the HIV env gene. Multiple computational approaches have been developed, offering a faster and more accessible alternative.
- Position-Specific Scoring Matrices (PSSM): This method uses two coreceptor-specific weight matrices (for R5 and X4) derived from large sequence databases. A new V3 sequence is scored against both matrices, and the tropism is predicted based on which matrix it more closely matches [72].
- Support Vector Machine (SVM)-based tools (e.g., Geno2Pheno): These machine learning tools classify sequences based on features like the V3 loop amino acid sequence, charge, and other physicochemical properties. They report a result along with a "false positive rate" (FPR), where a lower FPR indicates a higher probability of CXCR4 usage [71] [72].
- Hybrid and Next-Generation Methods: Newer methods aim to improve accuracy by incorporating information from the entire envelope gene, not just V3, and by using more advanced neural network computations [69] [74].
Table 2: Comparison of Coreceptor Tropism Determination Methods
| Method Type | Examples | Principle | Advantages | Disadvantages |
|---|---|---|---|---|
| Phenotypic | Trofile Assay | Directly tests virus entry using CCR5- or CXCR4-expressing cell lines | High accuracy, considered reference method | Slow, expensive, complex, low throughput |
| Genotypic | 11/25 Rule, PSSM, Geno2pheno | Computational prediction based on V3 loop sequence | Rapid, cheaper, high throughput, accessible | Historically lower sensitivity for detecting minor X4 populations; requires validation |
| Advanced Genotypic | Coreceptor-specific Weight Matrices (CM), Neural Networks | Incorporates full env sequence, charge rules, machine learning | Improved accuracy (e.g., 95% accuracy, 0.885 MCC reported for CM [72]) | Computational complexity, ongoing development |
The workflow for a typical genotypic tropism test, from sample collection to result, is outlined below.
The Research Toolkit: Essential Reagents and Assays
Table 3: Key Research Reagent Solutions for Coreceptor Tropism Research
| Reagent / Assay | Function / Application | Key Examples & Notes |
|---|---|---|
| Cell Line-Based Phenotypic Assays | In vitro determination of coreceptor usage by live virus. | MT-2, Ghost, U87-MAGI cell lines; Commercial Trofile assay [72]. |
| Coreceptor Inhibitors | To block specific entry pathways and confirm tropism; therapeutic development. | Maraviroc (CCR5 antagonist, FDA-approved); Plerixafor (AMD3100) (CXCR4 antagonist, not FDA for HIV) [69] [70]. |
| Genotypic Prediction Algorithms & Web Servers | In silico prediction of tropism from V3 sequence data. | Geno2pheno, WebPSSM; Custom coreceptor-specific matrices (CM) [71] [72]. |
| Monoclonal Antibodies for Flow Cytometry | Quantifying coreceptor expression and T-cell immune activation. | Anti-CCR5, Anti-CXCR4; Anti-HLA-DR, Anti-CD38 for activation markers [71]. |
| Humanized Mouse Models (in vivo Outgrowth Assay) | Ultra-sensitive detection of replication-competent virus in patient samples. | Used to validate cure after CCR5Δ32/Δ32 HSCT [75]. |
Therapeutic Implications and Future Directions
CCR5 Inhibitors and the Challenge of Resistance
The discovery of the CCR5Δ32 mutation paved the way for the development of CCR5 inhibitors, a class of entry inhibitors. Maraviroc is the prime example, licensed for clinical use in 2007 [69]. Its administration requires prior tropism testing to confirm the absence of CXCR4-using viruses. A significant challenge is the potential for virological failure during Maraviroc therapy, which, in over 50% of cases, involves the outgrowth of pre-existing, undetected X4 or dual-tropic viruses rather than the development of true resistance mutations against the drug itself [69]. In some cases, resistance does develop via envelope mutations that allow the virus to use the drug-bound CCR5 receptor, though these mutations often confer a fitness cost [69].
The Promise of Dual and Broad-Spectrum Antagonists
Given the virus's ability to switch tropism to evade single-receptor inhibitors, there is considerable interest in developing dual CCR5/CXCR4 antagonists [70] [74]. These compounds could prevent the escape of X4 variants and potentially be used in broader patient populations without the need for extensive tropism screening. Early examples include the peptide KR21 and small molecules like AMD3451 and certain pyrazole-based compounds [70]. The development of such multi-target inhibitors represents a promising next generation in entry inhibitor therapy.
CCR5Δ32/Δ32 Hematopoietic Stem Cell Transplantation: A Proof-of-Concept for Cure
The most profound demonstration of CCR5's critical role is the successful cure of HIV-1 in several individuals following allogeneic hematopoietic stem cell transplantation (HSCT) from donors homozygous for the CCR5Δ32 mutation [75]. This procedure effectively replaces the patient's immune system with one that is genetically resistant to R5-tropic HIV infection. Detailed virological and immunological follow-up of one such patient showed no replication-competent virus detected despite sophisticated assays, and no viral rebound occurred for four years after antiretroviral treatment interruption, providing strong evidence for a cure [75]. While HSCT is not a scalable solution for the millions living with HIV due to its high risk and cost, it validates CCR5 as a target for gene therapy approaches aimed at mimicking this protective effect.
The emergence of CXCR4-using HIV strains through coreceptor tropism switching remains a pivotal event in HIV pathogenesis, driving accelerated disease progression and complicating treatment. Research into the CCR5 coreceptor and the protective CCR5Δ32 mutation has been instrumental in shaping our understanding of viral entry and host-virus interactions. It has directly led to novel therapeutic classes, with the potential for even more robust dual antagonists on the horizon. Furthermore, it has provided a clear proof-of-concept that HIV can be cured through targeted genetic modification of the CCR5 gene. Ongoing research must continue to refine tropism prediction tools, elucidate the precise immunological triggers for the switch, and translate these insights into next-generation therapies and scalable cure strategies that can benefit a global population.
The CRISPR/Cas9 system has revolutionized genetic engineering, offering unprecedented precision in genome editing. In therapeutic contexts, particularly in the pursuit of an HIV cure, its application has shown remarkable promise. The primary therapeutic strategy involves disrupting the CCR5 gene, a coreceptor essential for HIV cellular entry [76]. Individuals carrying a natural 32-base pair deletion in the CCR5 gene (CCR5-Δ32) are resistant to HIV-1 infection, a phenomenon demonstrated by the curative outcomes of hematopoietic stem cell transplantations from CCR5Δ32/Δ32 donors [76] [18] [20]. CRISPR/Cas9 offers the potential to replicate this protective mutation in a patient's own cells, thereby creating a renewable source of HIV-resistant immune cells [53].
However, the efficacy and safety of this approach are critically dependent on the precision of the editing system. Off-target effects—unintended cuts at genomic sites with sequences similar to the intended target—pose a significant risk [77] [78]. These spurious edits can lead to mutations that disrupt tumor suppressor genes, activate oncogenes, or cause large-scale genomic rearrangements, potentially leading to adverse outcomes like malignant transformation [77] [79]. For therapies involving hematopoietic stem cells, which have the capacity for long-term self-renewal, the risk of propagating such mutations is of paramount concern. This technical guide provides an in-depth analysis of the mechanisms, detection methodologies, and minimization strategies for off-target effects, specifically within the context of developing a CRISPR-based cure for HIV.
Mechanisms of CRISPR/Cas9 Off-Target Effects
Understanding the molecular underpinnings of off-target activity is the first step toward its mitigation. The CRISPR/Cas9 system's specificity is governed by the interaction between the single-guide RNA (sgRNA) and the target DNA sequence, facilitated by the Cas9 nuclease. Several factors can compromise this specificity.
- sgRNA-DNA Mismatch Tolerance: The Cas9 nuclease can tolerate a limited number of mismatches—non-complementary base pairs—between the sgRNA and the genomic DNA, particularly if these mismatches are located distal to the Protospacer Adjacent Motif (PAM) sequence [77] [79]. The PAM (5'-NGG-3' for the standard Streptococcus pyogenes Cas9) is an essential recognition site for Cas9 activation. Research indicates that Cas9 can still cleave DNA even with up to three to five mismatches, depending on their position and distribution [77].
- The Seed Sequence: The 8-12 nucleotides closest to the PAM sequence, known as the "seed" region, are critical for target recognition. Mismatches within this region are less tolerated and generally reduce cleavage efficiency, whereas mismatches in the distal region are more easily overlooked by the Cas9 complex [79].
- Genomic and Structural Factors: Regions of the genome with high sequence homology to the intended target are natural hotspots for off-target activity [79]. Furthermore, the chromatin state and epigenetic modifications can influence accessibility; open chromatin regions are generally more susceptible to both on- and off-target cleavage than tightly packed heterochromatin [77] [79]. The GC content of the target site also plays a role, with an optimal range of 40-60% being associated with higher specificity [79].
The diagram below illustrates the primary molecular mechanisms that lead to off-target effects in the CRISPR/Cas9 system.
Detection and Prediction of Off-Target Effects
Accurately identifying off-target sites is a critical component of preclinical safety assessment. The methodologies for detection can be broadly classified into in silico prediction tools and experimental assays, each with distinct advantages and limitations.
In Silico Prediction Tools
Computational tools leverage algorithms to scan reference genomes for sequences with high similarity to the sgRNA, nominating potential off-target sites for further empirical validation.
Table 1: Comparison of In Silico Off-Target Prediction Tools
| Tool Name | Algorithm Type | Key Features | Key Limitations |
|---|---|---|---|
| CasOT [77] | Alignment-based | Exhaustive search; customizable PAM and mismatch parameters. | Does not account for chromatin environment. |
| Cas-OFFinder [77] | Alignment-based | High tolerance for various PAM types, mismatches, and bulges. | Purely sequence-based; results require validation. |
| CCTop [77] | Scoring-based | Scores based on distance of mismatches from the PAM sequence. | Limited by the accuracy of its scoring model. |
| DeepCRISPR [77] | Machine Learning | Incorporates both sequence and epigenetic features for prediction. | Requires substantial computational resources. |
While invaluable for initial sgRNA design and screening, in silico tools are inherently limited by their reliance on sequence data alone and may miss off-target sites influenced by the cellular milieu [77].
Experimental Detection Methods
Cell-based and cell-free experimental assays provide a more direct and comprehensive profile of off-target activity.
Table 2: Experimental Methods for Detecting Off-Target Effects
| Method | Principle | Advantages | Disadvantages |
|---|---|---|---|
| GUIDE-seq [77] | Captures double-stranded breaks (DSBs) via integration of double-stranded oligodeoxynucleotides. | High sensitivity; low false-positive rate; works in living cells. | Limited by transfection efficiency. |
| CIRCLE-seq [77] [79] | In vitro method using circularized, sheared genomic DNA incubated with Cas9/sgRNA. | Highly sensitive; low background; does not require living cells. | Performed in a cell-free system, may not reflect in vivo conditions. |
| Digenome-seq [77] [79] | Cas9/sgRNA-digested genomic DNA is subjected to whole-genome sequencing. | Highly sensitive; no transfection needed. | Expensive; requires high sequencing coverage. |
| SITE-Seq [77] [79] | Biochemical method with selective enrichment of cleaved fragments. | Minimal read depth; eliminates background noise. | Lower sensitivity and validation rate compared to other methods. |
| Whole Genome Sequencing (WGS) [77] [79] | Sequencing the entire genome of edited and control cells to identify mutations. | Comprehensive and unbiased analysis of the whole genome. | Very expensive; may miss low-frequency events in a heterogeneous cell population. |
The following workflow outlines a recommended, multi-layered strategy for a comprehensive off-target assessment, integrating both computational and empirical approaches.
Strategies for Minimizing Off-Target Effects
Multiple strategies have been developed to enhance the precision of CRISPR/Cas9 editing, focusing on optimizing the system's components and its delivery.
sgRNA Optimization: The design of the sgRNA is the most critical factor for specificity. Strategies include:
- Truncated sgRNAs: Shortening the sgRNA sequence by 2-3 nucleotides at the 5' end reduces its binding energy and increases its sensitivity to mismatches, thereby enhancing specificity [79].
- Chemical Modifications: Incorporating specific chemical modifications into the sgRNA backbone can improve its stability and precision, though this is an area of ongoing research [79].
- Bioinformatic Screening: Using multiple in silico tools to select sgRNAs with minimal homology to other genomic regions, optimal GC content (40-60%), and no predicted off-target sites in exonic or crucial regulatory regions [77] [79].
High-Fidelity Cas9 Variants: Wild-type Cas9 has been engineered to create high-fidelity mutants with reduced off-target activity. These variants, such as eSpCas9(1.1) and SpCas9-HF1, have altered amino acids that weaken the binding affinity to non-target DNA, making them less tolerant of mismatches while largely retaining on-target efficiency [77] [79].
Temporal Control of Cas9 Activity: The duration of Cas9 expression inside the cell is directly correlated with the risk of off-target effects. Using transient delivery methods, such as pre-assembled Cas9 protein-sgRNA ribonucleoprotein (RNP) complexes, limits the window of nuclease activity to a few hours, significantly reducing off-target edits compared to plasmid DNA transfection [53] [79]. This RNP approach was successfully used in a clinical trial (NCT03164135) for CCR5 knockout in hematopoietic stem cells, where edited cells persisted for over 19 months without detected adverse events [53].
A Case Study in HIV Research: CCR5 Gene Editing
The application of CRISPR/Cas9 to disrupt the CCR5 locus provides a compelling real-world example of off-target risk assessment and management. A 2024 study combined CCR5 knockout with a C46 HIV-1 fusion inhibitor to achieve resistance against both R5- and X4-tropic HIV strains [53].
Experimental Protocol:
- Cell Line: The MT4CCR5 cell line was used.
- Gene Editing: Cells were nucleofected with a pre-assembled RNP complex containing two sgRNAs targeting the first exon of the CCR5 gene (at the Δ32 mutation site) and in-house purified Cas9 protein.
- Dosage: Two RNP doses were tested (6μg Cas9 + 2μg of each sgRNA vs. 10μg Cas9 + 4μg of each sgRNA).
- Efficiency Assessment: Cleavage efficiency was confirmed via the T7 Endonuclease I (T7E1) assay, and CCR5 knockout was quantified by flow cytometry and Western blot. The higher RNP dose achieved a 97.89% reduction in CCR5 expression [53].
- Off-Target Assessment: The sgRNAs were specifically screened and selected for high cleavage efficiency and low off-target potential prior to the experiment, a crucial safety step [53].
The Scientist's Toolkit: Table 3: Essential Reagents for CRISPR-Mediated CCR5 Knockout
Research Reagent Function in the Experiment Recombinant Cas9 Protein The DNA endonuclease that creates double-strand breaks at the target site. sgRNAs targeting CCR5 Exon 1 Guides the Cas9 protein to the specific genomic locus for disruption. Nucleofection System Enables efficient delivery of the RNP complex into hard-to-transfect cells. T7 Endonuclease I Assay Kit Detects and quantifies the efficiency of genome editing by identifying mismatches in heteroduplex DNA. Flow Cytometry Antibodies Fluorescently-labeled antibodies against CCR5 to measure knockout efficiency at the protein level.
The journey toward a safe and effective CRISPR/Cas9-based therapy, particularly for a disease as complex as HIV, is intrinsically linked to solving the challenge of off-target effects. As demonstrated in the CCR5 editing paradigm, a multi-faceted approach is essential. This involves the careful selection and optimization of sgRNAs, the use of high-fidelity Cas9 variants, and the adoption of transient delivery methods like RNP complexes. Furthermore, a rigorous and multi-tiered off-target assessment strategy—combining sophisticated in silico prediction with sensitive empirical detection methods—is non-negotiable for preclinical safety profiling. While challenges remain, the continued refinement of these tools and strategies significantly de-risks the translational path. The ongoing clinical work to create CCR5-modified hematopoietic stem cells is a testament to this progress, bringing the goal of a broadly applicable CRISPR-mediated HIV cure closer to reality.
The C-C chemokine receptor 5 (CCR5) is a G-protein coupled receptor (GPCR) that plays a critical role in immune cell migration and inflammatory responses [17] [2]. While widely recognized as the primary coreceptor for human immunodeficiency virus (HIV) entry into target cells, CCR5 exhibits pleiotropic functions beyond viral pathogenesis, influencing susceptibility to various pathogens and contributing to multiple disease processes [17] [80]. The discovery that a natural 32-base pair deletion in the CCR5 gene (CCR5-Δ32) confers strong resistance to HIV-1 infection in homozygous individuals sparked intense research interest in this receptor as a therapeutic target [18] [20]. However, this protective effect against HIV must be balanced against emerging evidence that CCR5 deficiency may alter susceptibility to other infectious diseases, creating a complex landscape for therapeutic intervention [17] [80]. This review examines the dual nature of CCR5-mediated pathogen responses, highlighting the critical balance between HIV protection and potential vulnerabilities to other infections.
CCR5 Structure, Function, and Expression
Molecular Structure and Signaling Mechanisms
CCR5 belongs to the class A family of GPCRs and features a characteristic seven-transmembrane domain structure connected by three intracellular and three extracellular loops [17] [2]. The receptor possesses two disulphide bridges connecting the N-terminal domain to the third extracellular loop (ECL3) and transmembrane domain 3 to ECL2, which are critical for maintaining proper receptor conformation [2]. The natural ligands for CCR5 include chemokines CCL3 (MIP-1α), CCL4 (MIP-1β), CCL5 (RANTES), and CCL3L1, which bind to the receptor and initiate intracellular signaling cascades [17].
Upon ligand binding, CCR5 undergoes conformational changes that stimulate pertussis toxin-sensitive heterotrimeric G proteins, catalyzing the exchange of GTP for GDP in the Gα subunit [17]. This activation triggers multiple downstream signaling pathways, including:
- Chemotaxis and immune cell migration through MAPK and PI3K pathways
- Calcium flux and cell activation via phospholipase C activation
- Inflammatory gene expression through NF-κB and AKT pathways [17] [80]
Following activation, CCR5 rapidly undergoes phosphorylation in the carboxy-terminal region, promoting desensitization and internalization mediated by β-arrestin, which sequesters the receptor to clathrin-coated pits for endocytosis and subsequent recycling [17].
Tissue Expression and Physiological Roles
CCR5 is expressed on a wide array of bone-marrow-derived cells, including:
- Lymphocytes (T cells, particularly TH1 and CD8+ subsets)
- Myeloid cells (monocytes, macrophages, dendritic cells)
- Other immune cells (NK cells, regulatory T cells) [17]
The receptor is found in primary and secondary lymphoid organs (thymus, spleen), non-hematopoietic peripheral tissues (epithelium, endothelium, vascular smooth muscles, fibroblasts), and the central nervous system (neurons, astrocytes, microglia) [17]. In normal adult brain, CCR5 is highly expressed in microglia but undetectable in neurons, though neuronal expression can be induced under pathological conditions [17].
Table 1: CCR5 Expression Across Cell Types and Tissues
| Cell/Tissue Type | Expression Level | Primary Function |
|---|---|---|
| CD4+ T lymphocytes (TH1 subset) | High | T-cell activation and trafficking |
| Monocytes/Macrophages | High | Inflammatory response |
| Microglia | High | CNS immunosurveillance |
| Dendritic cells | Moderate | Antigen presentation |
| Neurons | Low/Inducible | Synaptic plasticity (pathological conditions) |
| Hepatic cells | Moderate | Inflammatory recruitment |
CCR5's primary physiological role involves directing immune cell migration (chemotaxis) along chemokine gradients to sites of inflammation and infection [17]. Beyond its immunoregulatory functions, CCR5 also suppresses learning, memory formation, and synaptic connections in the brain, suggesting important neuromodulatory activities [17] [20].
The CCR5-Δ32 Mutation: Molecular Mechanisms and HIV Resistance
Genetic Basis and Global Distribution
The CCR5-Δ32 variant is characterized by a 32-base-pair deletion in the CCR5 gene coding region that introduces a premature stop codon, resulting in a truncated, non-functional receptor that fails to reach the cell surface [20]. In homozygous individuals (Δ32/Δ32), functional CCR5 receptors are absent from the cell surface, conferring high-level resistance to CCR5-tropic HIV-1 strains [18] [20]. Heterozygous individuals (+/Δ32) exhibit approximately 50% reduction in surface CCR5 expression due to dimerization between mutant and wild-type receptors that interferes with proper receptor trafficking [20].
The global distribution of CCR5-Δ32 shows striking geographical patterns:
- Highest frequency: Northern European populations (up to 16% in Finland and Russia)
- Moderate frequency: Other European populations (10-14% average)
- Virtual absence: African, East Asian, and Native American populations [17] [20]
This distribution suggests a single mutation event occurring in Northern European populations approximately 700-2,500 years ago, with subsequent spread possibly facilitated by Viking migrations [17] [20].
Protective Mechanisms Against HIV Infection
The CCR5-Δ32 mutation confers HIV resistance through multiple mechanisms:
- Loss of viral entry point: The truncated receptor cannot localize to the cell surface, preventing HIV gp120 from binding and initiating fusion [20]
- Dominant negative effect: In heterozygotes, mutant-wild type receptor dimerization reduces surface expression of functional CCR5 [20]
- Altered immune environment: Reduced CCR5 signaling may create less favorable conditions for viral replication [17]
Table 2: HIV Susceptibility by CCR5 Genotype
| Genotype | Cell Surface CCR5 | HIV Susceptibility | Disease Progression |
|---|---|---|---|
| Wild type (+/+) | Normal | High | Standard progression |
| Heterozygous (+/Δ32) | ~50% reduction | Reduced | 2-3 year delay in AIDS onset |
| Homozygous (Δ32/Δ32) | Absent | Highly resistant | Protection from infection |
Meta-analyses of case-control studies confirm that CCR5-Δ32 homozygosity significantly reduces HIV-1 susceptibility (OR=0.25, 95%CI=0.09-0.68), while heterozygosity may slightly increase susceptibility (OR=1.16, 95%CI=1.02-1.32) but delays disease progression in infected individuals [12].
CCR5 in Non-HIV Pathogenesis: Beyond Viral Entry
Altered Susceptibility to Other Pathogens
The pleiotropic nature of CCR5 means that its deletion or inhibition influences susceptibility to various pathogens beyond HIV:
Viral Infections:
- West Nile Virus: Δ32 homozygotes show increased susceptibility to symptomatic West Nile virus infection, with more severe neurological manifestations [80]
- Influenza: CCR5 knockout mice demonstrate impaired response to Influenza A infection, suggesting CCR5 is important for efficient antiviral response [20]
- Tick-borne encephalitis: CCR5 deficiency may increase risk of severe disease progression [80]
Bacterial Infections:
- Mycobacterium tuberculosis: CCR5 mediates recruitment of protective T-cell responses; its absence may impair control of tuberculosis [17]
- Yersinia pestis: Despite historical theories linking Δ32 to plague resistance, mouse studies show no protective effects [20]
Parasitic Infections:
- Plasmodium species: Altered immune responses in CCR5 deficiency may affect malaria pathogenesis, though data remains limited [17]
The diagram below illustrates the dual nature of CCR5 in pathogen defense:
Evolutionary Pressures and Historical Pathogens
The high frequency of CCR5-Δ32 in European populations despite its detrimental effects against some pathogens suggests strong historical selective pressure. Competing theories propose different driving pathogens:
Smallpox Hypothesis:
- Stronger scientific support links Δ32 to smallpox (Variola major) protection [20]
- Smallpox has sufficient historical depth (2000+ years) to account for Δ32 frequency
- Myxoma virus (a smallpox relative) uses CCR5 for entry, supporting mechanistic plausibility [20]
- Higher smallpox mortality in children creates stronger selective pressure [20]
Plague Hypothesis:
- Originally proposed due to timing coincidence with Black Death (1346-1352) [20]
- Lacks supporting evidence from mouse infection models [20]
- Yersinia pestis (plague bacterium) biologically distinct from viruses that typically use CCR5 [20]
The evolutionary trajectory of CCR5-Δ32 demonstrates the complex balance of genetic advantages, where protection against one devastating pathogen (smallpox) may come at the cost of increased susceptibility to others (West Nile virus) [17] [20].
Experimental Approaches and Research Methods
Coreceptor Function Assays
Viral Entry Assays
- Methodology: Utilize pseudotyped viruses expressing HIV-1 envelope proteins in combination with CD4 and CCR5/CXCR4-expressing cell lines (e.g., U87.CD4.CCR5/CXCR4, HEK293T) [2]
- Key measurements: Luciferase reporter activity, p24 antigen production, GFP expression
- Applications: Determine coreceptor tropism, evaluate entry inhibitors, assess impact of CCR5 mutations [2]
Calcium Flux Measurements
- Procedure: Load CCR5-expressing cells with calcium-sensitive dyes (Fura-2, Fluo-4), stimulate with chemokine ligands (CCL3, CCL4, CCL5) or gp120, monitor intracellular calcium changes via fluorometry [81]
- Outcome: Assess G-protein coupling and receptor activation efficiency [81]
Gene Editing Techniques
CRISPR-Cas9 Approaches
- Protocol: Design sgRNAs targeting CCR5 exon regions, deliver via lentiviral vectors to primary CD4+ T-cells or hematopoietic stem cells, validate editing efficiency via T7E1 assay or sequencing [18]
- Applications: Create CCR5-knockout cells for functional studies, develop therapeutic approaches for HIV resistance [18]
Zinc Finger Nuclease Methods
- Procedure: Engineer sequence-specific zinc finger proteins coupled to nuclease domains, electroporate into target cells, select modified populations via FACS or drug selection [18]
- Clinical relevance: Basis for SB-728-T clinical trials for HIV functional cure [18]
Structural Biology Techniques
X-ray Crystallography
- Methodology: Express and purify stabilized CCR5 constructs, crystallize in complex with ligands (maraviroc, chemokines), solve structure via molecular replacement [2]
- Key findings: Revealed CCR5 conformational states and maraviroc binding site in extracellular vestibule [2]
Nuclear Magnetic Resonance (NMR)
- Approach: Label CCR5 with stable isotopes (15N, 13C), collect spectra in micellar solutions, determine structure constraints from chemical shifts and NOEs [2]
- Advantage: Studies receptor dynamics and ligand interactions in solution state [2]
The experimental workflow for comprehensive CCR5 investigation is shown below:
Research Reagent Solutions
Table 3: Essential Research Tools for CCR5 Investigation
| Reagent Category | Specific Examples | Research Applications |
|---|---|---|
| Cell Lines | U87.CD4.CCR5, HEK293T-CCR5, PM1-CCR5 | Viral entry assays, receptor function studies |
| Antibodies | Anti-CCR5 (clone 2D7), Anti-CD4, Anti-CCR5-phycoerythrin | Flow cytometry, receptor quantification, blocking studies |
| Chemical Inhibitors | Maraviroc, Cenicriviroc, TAK-779 | Coreceptor blockade, signaling inhibition studies |
| Recombinant Proteins | CCL3, CCL4, CCL5, gp120 | Ligand binding assays, signaling activation, competition studies |
| Gene Editing Tools | CRISPR-Cas9 constructs, ZFNs, shRNAs | CCR5 knockout/knockdown, therapeutic development |
| Animal Models | CCR5-/- mice, Humanized mouse models | In vivo pathogenesis studies, therapeutic efficacy testing |
Therapeutic Implications and Future Directions
CCR5-Targeted HIV Interventions
The development of CCR5-targeted therapies represents a promising approach for HIV treatment and prevention:
Small Molecule Inhibitors:
- Maraviroc: Only FDA-approved CCR5 antagonist, acts as allosteric inhibitor that stabilizes CCR5 in inactive conformation [2]
- Clinical application: Used in combination antiretroviral therapy for R5-tropic HIV [2]
Gene Therapy Approaches:
- Sangamo's SB-728-T: Zinc finger nuclease-modified CD4+ T-cells showing reduced viral load in clinical trials [18]
- CRISPR-Cas9 strategies: Preclinical development of CCR5-edited hematopoietic stem cells for HIV functional cure [18]
Stem Cell Transplantation:
- Berlin Patient: First documented HIV cure following transplantation with CCR5-Δ32/Δ32 hematopoietic stem cells [18]
- London Patient: Second confirmed case of HIV remission following similar approach [18]
Balancing Therapeutic Efficacy with Safety Concerns
Therapeutic CCR5 targeting requires careful consideration of potential consequences:
Neurological Effects:
- CCR5 suppression enhances learning and memory in animal models [20]
- CCR5 over-activation by viral proteins may contribute to HIV-associated cognitive deficits [20]
- Therapeutic inhibition might produce unexpected neurological benefits [20]
Inflammatory and Autoimmune Considerations:
- CCR5 blockade shows benefit in reducing liver fibrosis in nonalcoholic steatohepatitis [2]
- CCR5 inhibition may have applications in cancer immunotherapy by enhancing T-cell infiltration [2]
- Potential exists for increased susceptibility to specific pathogens (West Nile virus, influenza) [80]
Future Research Priorities
Key areas for future investigation include:
- Tissue-specific CCR5 modulation: Developing approaches that target CCR5 in specific cell types or tissues to minimize systemic effects
- Dual receptor targeting: Exploring combination approaches that address both CCR5 and alternative pathways
- Personalized risk assessment: Developing biomarkers to identify individuals who would benefit most from CCR5-targeted therapies while minimizing risks
- Long-term safety studies: Comprehensive evaluation of CCR5 inhibition in diverse patient populations
CCR5 represents a paradigm of balanced polymorphism in human genetics, where a single receptor influences susceptibility to multiple pathogens through its pleiotropic functions. The CCR5-Δ32 mutation provides remarkable protection against HIV infection but simultaneously alters susceptibility to other infectious agents, particularly increasing risk for symptomatic West Nile virus infection. This balance between protective and detrimental effects illustrates the complex evolutionary trade-offs that shape human immune genetics. Future therapeutic strategies targeting CCR5 must carefully consider this delicate balance, developing approaches that maximize benefits while minimizing potential risks associated with altered pathogen susceptibility. As gene editing technologies advance toward clinical application, comprehensive understanding of CCR5's diverse roles will be essential for developing safe, effective interventions that harness its protective potential without unintended consequences.
The pursuit of an HIV cure is fundamentally challenged by profound clinical heterogeneity, encompassing diverse host genetics, variable viral reservoirs, and divergent patient-specific disease progression. The seminal cases of the "Berlin" and "London" patients, who achieved sustained HIV remission following hematopoietic stem cell transplantation from donors with a homozygous CCR5Δ32 mutation, established the foundational principle that genetic disruption of the HIV co-receptor CCR5 can confer natural resistance to R5-tropic HIV strains [3] [82]. This breakthrough illuminated a definitive path toward a functional cure while simultaneously revealing a landscape of extreme complexity. The CCR5 pathway, critical for macrophage and T-cell infection by M-tropic HIV strains, represents an ideal therapeutic target; individuals lacking functional CCR5 demonstrate significant resistance to HIV infection without apparent pathological consequences, making this receptor a compelling focus for pharmacological and genetic intervention [5]. However, the translational application of this knowledge is complicated by several layers of heterogeneity: variable CCR5Δ32 allele frequencies across ethnic populations [82], the emergence of CXCR4-tropic viruses following CCR5 blockade [3], and a remarkably diverse latent reservoir that persists despite antiretroviral therapy (ART) [83] [84]. This whitepaper delineates a personalized framework to navigate this multidimensional heterogeneity, integrating advanced genomic tools, single-cell reservoir characterization, and synergistic therapeutic strategies to architect the next generation of HIV cure paradigms.
Host Genetic Heterogeneity: The CCR5Δ32 Distribution Landscape
The natural CCR5Δ32 mutation, a 32-base-pair deletion rendering the receptor non-functional, demonstrates significant geographical and ethnic variation, directly influencing the feasibility of donor-dependent therapies and population-wide treatment strategies.
Table 1: Global Distribution of CCR5Δ32 Allele Frequencies [82]
| Population | Sample Size | Δ32 Allele Frequency (%) | Notes |
|---|---|---|---|
| Northern European | 8,442 (across 40 studies) | 7.7 - 10.0 | Highest frequencies observed |
| North American Caucasians | Not Specified | ~21.7% Heterozygosity | Derived primarily from European ancestry |
| African Populations | Not Specified | 0% | Allele largely absent |
| Japanese Population | Not Specified | 0% | Allele largely absent |
| African-Americans | Not Specified | 5.8% Heterozygosity | Reflecting genetic admixture |
| Hispanic-Americans | Not Specified | 6.9% Heterozygosity | Reflecting genetic admixture |
| Asian-Americans | Not Specified | 0.6% Heterozygosity | Very rare |
This heterogeneous genetic distribution necessitates personalized screening protocols prior to considering any CCR5-targeted therapeutic intervention. The observed north-to-south cline of decreasing Δ32 frequency within Europe further underscores the need for geographically tailored approaches [82]. For populations with low or null Δ32 frequency, alternative strategies such as autologous cell therapy using CRISPR/Cas9 to engineer the Δ32 mutation in a patient's own cells become critically important [85].
Experimental Protocol: CCR5Δ32 Genotyping and Quantification
Objective: To accurately identify the CCR5Δ32 genotype and quantify the frequency of mutant alleles in heterogeneous cell mixtures, such as those following stem cell transplantation or gene-editing procedures.
Methodology: Droplet Digital PCR (ddPCR) [85]
- DNA Extraction: Isolate genomic DNA from patient peripheral blood mononuclear cells (PBMCs) or hematopoietic stem/progenitor cells using a standard phenol-chloroform method or commercial kit.
- Primer/Probe Design: Design two primer-probe sets:
- Wild-type CCR5 Probe: Fluorescently labeled (e.g., FAM), targeting the intact 32bp sequence.
- Mutant CCR5Δ32 Probe: Fluorescently labeled with a different fluorophore (e.g., HEX/VIC), specifically spanning the deletion junction.
- Reaction Setup: Partition the PCR reaction mixture, containing the target DNA and both probe sets, into approximately 20,000 nanoliter-sized droplets.
- Endpoint PCR: Amplify the target sequences using optimized thermal cycling conditions.
- Droplet Reading and Analysis: Analyze each droplet individually using a droplet reader. Count the number of positive droplets for FAM (wild-type), HEX/VIC (Δ32), or both (heterozygous), and apply Poisson statistics to calculate the absolute copy number and frequency of each allele in the original sample.
This method provides superior quantification down to a sensitivity of 0.8% mutant alleles in a mixed population, enabling precise monitoring of engraftment success for gene-edited or donor-derived cells [85].
Viral and Reservoir Heterogeneity: A Multi-Faceted Obstacle
Beyond host genetics, heterogeneity within the virus and the latent reservoir presents a formidable barrier to cure.
Viral Tropism and Coreceptor Switching
HIV entry requires binding to both the CD4 receptor and a coreceptor, predominantly CCR5 (R5-tropic) or CXCR4 (X4-tropic). R5-tropic viruses dominate early infection, but a significant clinical challenge is viral tropism switching. Under selective pressure from CCR5-targeted therapies, the virus may evolve to utilize CXCR4, enabling continued infection and pathogenesis [3] [5]. This necessitates dual-targeted intervention strategies.
The Heterogeneous Latent Reservoir
The HIV latent reservoir, primarily composed of resting memory CD4+ T cells, is not a monolithic entity but a highly diverse collection of infected cells. Single-cell multi-omics has revealed its profound phenotypic and epigenetic heterogeneity [83] [84].
Table 2: Heterogeneity of the HIV-1 Latent Reservoir in CD4+ T Cells [83] [84]
| Reservoir Characteristic | Subtypes and Markers | Functional Significance |
|---|---|---|
| Memory CD4+ T Cell Subsets | Naive (TN), Central Memory (TCM), Transitional Memory (TTTM), Effector Memory (TEM), Terminally Differentiated (TTD) | TCM cells are long-lived and abundant, forming a primary reservoir. TEM have higher proliferation rates. |
| CD4+ T Helper Cell Subsets | Th1, Th2, Th17, Follicular Helper T (TFH), Regulatory T (Treg) | TFH cells are major reservoirs in lymphoid tissues. Th1/Th17 cells are long-term reservoirs. Treg cells harbor frequent provirus and pose an immune evasion challenge. |
| Surface Markers Enriched on HIV+ Cells | CCR5, SLAM, PD-1, CD2, CD25, CD95, ICOS [83] | Identifies cells prone to infection and persistence; provides targets for reservoir enrichment and elimination. |
| Non-T Cell Reservoirs | Macrophages, Microglia, Astrocytes | Long-lived, tissue-resident, and resistant to HIV's cytopathic effects; contribute to anatomical reservoir complexity. |
The following diagram illustrates the complex cellular heterogeneity of the latent HIV reservoir and the key surface markers identified on infected cells.
Experimental Protocol: Single-Cell Reservoir Characterization via ASAP-seq
Objective: To precisely define the unperturbed HIV-infected cell reservoir from ART-treated patients by simultaneously detecting integrated proviral DNA, epigenetic state, and cell surface protein profiles at single-cell resolution [83].
Methodology: Assay for Transposase-Accessible Chromatin with Select Antigen Profiling (ASAP-seq) [83]
- Sample Preparation: Isolate PBMCs from ART-treated people living with HIV (ART-PLWH). Include controls from uninfected donors.
- Antibody Staining: Label cells with a panel of metal-tagged antibodies targeting surface proteins of interest (e.g., CD4, CCR5, PD-1, CD2).
- Nuclei Isolation and Transposition: Permeabilize cells and isolate nuclei. Use the Tn5 transposase to fragment and tag accessible genomic regions with sequencing adapters.
- Library Preparation and Sequencing: Generate separate sequencing libraries for the chromatin accessibility data (ATAC-seq) and the antibody-derived tags (ADT).
- Bioinformatic Analysis:
- Alignment: Map sequencing reads to a combined human (hg38) and HIV (e.g., SUMA strain) reference genome.
- Cell Calling: Identify cells containing reads that align to integrated HIV provirus (HIV+ cells).
- Multi-omic Integration: Cluster cells based on epigenetic profiles and overlay surface protein expression and HIV status to define the phenotypic identity of HIV+ cells.
This protocol directly identifies infected cells without prior stimulation, revealing their native epigenetic state and surface signature, which is critical for designing targeted elimination strategies [83].
A Personalized Framework for HIV Cure Strategies
To address the outlined heterogeneities, a multi-pronged, personalized framework is essential. The core strategy involves synergistic multi-target gene editing combined with immunotherapy, tailored through patient-specific profiling.
Synergistic Multi-Target Gene Editing
A single-target approach against CCR5 is insufficient due to the risk of CXCR4 tropism switching and latent virus reactivation. A comprehensive viral blockade requires multiplexed gene editing [3].
- CCR5 Disruption: Renders primary target cells resistant to the most common R5-tropic strains. Techniques include CRISPR/Cas9, ZFNs, and TALENs to induce the Δ32-like knockout [3] [85].
- CXCR4 Disruption: A safeguard against coreceptor switching, providing broad resistance against both R5- and X4-tropic viruses.
- HIV LTR Targeting: Directly disrupts the viral promoter within the integrated provirus in latently infected cells, preventing viral reactivation and replication, or even excising the provirus [3].
The following diagram illustrates this multi-target gene editing strategy and its functional outcome in creating resistant cells.
Integrating Gene Editing with Immunotherapy
Gene-edited, HIV-resistant cells can be further empowered as effector vehicles for cure strategies.
- CAR-T Cells: Engineer CD4+ T cells or hematopoietic stem cells with both a CCR5Δ32 knockout and an HIV-specific Chimeric Antigen Receptor (CAR). These dual-function cells are resistant to infection and capable of targeting and killing HIV-infected cells presenting viral antigens [3].
- Immune Checkpoint Modulation: HIV-specific T cells often become "exhausted," expressing markers like PD-1. Combining gene editing with PD-1 blockade (e.g., using anti-PD-1 antibodies or secreting scFv fragments) can rejuvenate these cells, enhancing their capacity to clear the reservoir [3] [83].
The Scientist's Toolkit: Essential Research Reagents and Methodologies
Table 3: Key Research Reagent Solutions for HIV Reservoir and Cure Studies
| Reagent / Tool | Function / Application | Example Use Case |
|---|---|---|
| Droplet Digital PCR (ddPCR) | Absolute quantification of specific DNA sequences without a standard curve. | Precise measurement of CCR5Δ32 allele frequency in cell mixtures post-transplant/gene-editing [85]. |
| CRISPR/Cas9 System | Precise genome editing via a guide RNA (gRNA) and Cas9 nuclease. | Introduction of CCR5Δ32 mutation into wild-type CD4+ T cells or hematopoietic stem cells [3] [85]. |
| Single-Cell Multi-omics (ASAP-seq) | Simultaneous profiling of chromatin accessibility, surface protein expression, and integrated provirus in single cells. | Unbiased identification and phenotypic/epigenetic characterization of the latent HIV reservoir in patient samples [83]. |
| Latency Reversal Agents (LRAs) | Compounds that reactivate transcription of latent HIV provirus. | Used in "Shock and Kill" strategies to expose latent reservoirs for immune-mediated clearance (e.g., HDAC inhibitors, PKC agonists) [84]. |
| HIV-1 Envelope GP120 & Chemokines (MIP-1α, MIP-1β, RANTES) | Natural ligands for CCR5; used to study receptor binding, signaling, and viral entry inhibition. | In vitro assays to validate CCR5 function post-editing or to test efficacy of CCR5 antagonists [5]. |
Addressing the intertwined challenges of host genetic and viral reservoir heterogeneity is paramount for advancing an HIV cure. The path forward lies in the development of personalized treatment frameworks that begin with deep patient profiling—genotyping for CCR5Δ32 and coreceptor tropism, and mapping the latent reservoir via advanced single-cell assays. This diagnostic foundation must then inform the application of synergistic, multi-targeted interventions, such as multiplex gene editing combined with enhanced immunotherapy. While significant hurdles in safety, delivery, and global accessibility remain, this integrative and precision-focused approach provides a robust roadmap for overcoming the persistent barrier of clinical heterogeneity and achieving a functional cure for HIV.
The discovery that the C-C chemokine receptor type 5 (CCR5) serves as a major co-receptor for human immunodeficiency virus (HIV) entry into CD4+ T cells marked a pivotal advancement in virology and immunology [3] [86]. This understanding was profoundly illuminated by individuals carrying a natural 32-base pair deletion (CCR5Δ32) in the CCR5 gene, which renders the receptor non-functional and confers significant resistance to infection by CCR5-tropic (R5-tropic) HIV strains, the variants most prevalent during early and chronic stages of infection [3] [86]. The therapeutic potential of exploiting this natural resistance was definitively demonstrated by the cases of the "Berlin Patient" and "London Patient," who achieved long-term HIV remission following allogeneic hematopoietic stem cell transplantation (allo-HSCT) from donors homozygous for the CCR5Δ32 mutation, performed to treat their underlying hematologic malignancies [55] [3]. These cases provide a compelling proof-of-concept that transplanting CCR5-disrupted hematopoietic stem and progenitor cells (HSPCs) can reconstitute an HIV-resistant immune system.
However, the broader application of this cure strategy is severely constrained by significant challenges in HSPC delivery and engraftment. Allogeneic transplants from matched CCR5Δ32/Δ32 donors are not a scalable solution due to the rarity of the mutation, the risks of graft-versus-host disease (GvHD), and the procedural morbidity itself [86] [87]. Consequently, the field is shifting towards autologous transplantation of genetically modified HSPCs, where a patient's own cells are engineered to disrupt the CCR5 gene ex vivo and then reinfused [87]. The success of this approach, and indeed any HSPC-based therapy, hinges on overcoming multifactorial obstacles related to the efficient delivery of editing tools, the achievement of high-level engraftment of modified cells, and the subsequent reconstitution of a durable, resistant immune system. This technical guide delves into these core challenges, framing them within the critical context of developing a scalable cure for HIV.
Core Challenges in HSPC Therapy Delivery and Engraftment
The pathway from HSPC collection to stable engraftment of engineered cells is fraught with technical hurdles that directly impact therapeutic efficacy. The following table summarizes the primary challenges and their specific impacts on the goal of HIV remission.
Table 1: Key Challenges in HSPC Therapy for HIV Cure
| Challenge Category | Specific Challenge | Impact on Therapeutic Outcome |
|---|---|---|
| Editing Efficiency | Achieving high rates of biallelic CCR5 disruption in HSPCs. | Low efficiency allows unedited cells to support viral rebound, as demonstrated by a trial where low editing rates failed to prevent recrudescence [87]. |
| Conditioning Regimen | Balancing myeloablation to create niche space with regimen-related toxicity. | Reduced-intensity conditioning may compromise engraftment levels, while intense regimens increase patient risk [55] [86]. |
| Donor Chimerism | Achieving and maintaining full donor chimerism in allogeneic settings. | Mixed chimerism leaves host-derived, HIV-susceptible cells that can reignite infection [55]. |
| Tropism Switching | Emergence of CXCR4-tropic (X4-tropic) HIV variants post-CCR5 disruption. | Renders CCR5 knockout ineffective, leading to viral rebound, as seen in the "Essen Patient" [3] [87]. |
| Reservoir Clearance | Eliminating the latent HIV reservoir in host tissues. | Residual infected cells can cause rebound if the new immune system is not fully protective [88]. |
| Immune Reconstitution | Timely and functional reconstitution of CD4+ T cells post-transplant. | Slow CD4+ T cell recovery prolongs immunodeficiency and delays assessment of cure [55]. |
The Engraftment Barrier and Conditioning
A foundational requirement for successful HSPC therapy is the creation of vacant niches in the bone marrow for the transplanted cells to home to and engraft. This is achieved through conditioning regimens, which involve chemotherapy and/or radiation. The intensity of this regimen is a critical determinant. One key lesson from the "London Patient" was that HIV-1 remission could be achieved with a single allo-HSCT and a reduced-intensity conditioning regimen that did not include total body irradiation, suggesting that excessively aggressive and toxic conditioning may not be necessary [55]. However, in autologous settings, insufficient conditioning can lead to poor engraftment of the edited HSPCs, as the resident, unedited HSPCs outcompete them for niche space. This directly limits the proportion of circulating immune cells that are resistant to HIV, creating a population vulnerable to infection and potential viral rebound.
The Challenge of Viral Tropism and Multi-Target Strategies
A significant limitation of strategies focusing solely on CCR5 disruption is the potential for viral escape through tropism switching. HIV can adapt to use an alternative co-receptor, CXCR4, and CXCR4-tropic viruses are present in a substantial minority (estimated 18%–52%) of people living with HIV [87]. The case of the "Essen Patient" demonstrated this risk, where transplantation of CCR5Δ32/Δ32 cells was followed by rapid viral rebound of a pre-existing CXCR4-tropic variant [55] [3]. This underscores that single-target approaches are inherently vulnerable. Consequently, the field is moving towards multiplex gene editing strategies designed to create a comprehensive viral barrier. These approaches aim to simultaneously disrupt both major co-receptors, CCR5 and CXCR4, and/or target the integrated HIV proviral DNA itself, for instance, via editing of the viral Long Terminal Repeat (LTR) region to prevent reactivation [3]. Such multi-layered defenses are likely essential for a broadly applicable cure.
Experimental Protocols and Methodological Frameworks
To address the challenges outlined above, researchers have developed sophisticated experimental workflows. The following diagram and table detail a protocol for a combinatorial gene-editing approach, which represents the cutting edge of HSPC therapy for HIV.
Diagram 1: Workflow for Combinatorial HSPC Therapy
Table 2: Research Reagent Solutions for Combinatorial HSPC Engineering
| Research Reagent / Tool | Function in Experimental Protocol |
|---|---|
| CRISPR/Cas9 System | Precision nuclease for creating targeted double-strand breaks in genomic DNA (e.g., at CCR5, CXCR4 loci) [3] [87]. |
| AAV6 Donor Template | Adeno-associated virus serotype 6, used as a vector to deliver homologous recombination donor DNA for precise gene knock-in [87]. |
| Broadly Neutralizing Antibody (bNAb) Genes | Genes encoding potent antibodies (e.g., 10-1074, PGDM1400) knocked into the CCR5 locus to enable endogenous secretion [87]. |
| CD34+ HSPC Isolation Kit | Magnetic-activated cell sorting (MACS) or fluorescence-activated cell sorting (FACS) reagents for purifying hematopoietic stem cells from apheresis product. |
| StemSpan SFEM Medium | Serum-free, cytokine-supplemented culture medium optimized for the maintenance and expansion of HSPCs ex vivo. |
| TZM-bl Reporter Cell Line | An engineered HeLa cell line expressing CD4, CCR5, and CXCR4, used for quantifying HIV neutralization by secreted bNAbs [87]. |
Protocol: Multilayered Engineering of HSPCs for HIV Resistance
This protocol is designed to achieve both cell-intrinsic and cell-extrinsic HIV resistance, building on a strategy validated in pre-clinical models [87].
- HSPC Mobilization and Collection: Mobilize HSPCs from a patient using granulocyte-colony stimulating factor (G-CSF). Collect cells via apheresis.
- CD34+ Cell Isolation and Culture: Isulate CD34+ HSPCs from the apheresis product using clinical-grade magnetic bead separation. Activate the cells in serum-free medium supplemented with cytokines (SCF, TPO, FTL-3 ligand).
- Multiplex CRISPR-Cas9 Editing:
- Electroporation: Introduce CRISPR-Cas9 ribonucleoproteins (RNPs) targeting the CCR5 locus and, if desired, the CXCR4 locus, via electroporation.
- Knock-in of bNAb Cassettes: Co-electroporate with an AAV6 donor vector containing a payload designed for integration into the CCR5 locus. This payload should contain expression cassettes for one or more broadly neutralizing antibodies (e.g., 10-1074, Ibalizumab), linked as a single transcript to ensure proper pairing [87].
- Quality Control and Potency Assay:
- Editing Efficiency: Assess the frequency of indels at the CCR5 locus using next-generation sequencing (NGS). The goal is to maximize the percentage of biallelic disruption.
- Antibody Secretion: Differentiate a portion of the engineered HSPCs into B cells in vitro. Collect the supernatant and use a TZM-bl neutralization assay to confirm the secretion of functional antibodies that can inhibit both R5-tropic and X4-tropic HIV pseudoviruses [87].
- Transplantation and Engraftment:
- Conditioning: The patient undergoes myeloablative conditioning (e.g., with busulfan) to create space in the bone marrow.
- Infusion: The engineered HSPC product is infused back into the patient.
- Monitoring: Track engraftment (neutrophil and platelet recovery), donor chimerism (in allogeneic settings), and the persistence of edited cells in peripheral blood over time.
Advanced Strategies and Future Directions
Synergizing Gene Editing and Immunotherapy
The integration of gene editing with immunotherapy represents a paradigm shift with the potential to overcome persistent barriers. Engineered HSPCs can give rise to immune cells that not only are resistant to infection but also possess enhanced abilities to clear the latent reservoir. For instance, researchers are developing chimeric antigen receptor (CAR) T cells derived from edited HSPCs that can target and eliminate HIV-infected cells [3]. Furthermore, the immunosuppressive bone marrow microenvironment post-transplant can be modulated. The unexpected sustained HIV remission in a patient ("the Geneva patient") who received a transplant from a wild-type CCR5 donor and was maintained on the JAK1/2 inhibitor Ruxolitinib for chronic GvHD suggests that this drug class may have a beneficial role. Ruxolitinib may potentially reduce reservoir activation by modulating immune activation, highlighting how adjunctive immunotherapies can synergize with transplant-based cure strategies [88].
Safety and Personalization
As these therapies advance, long-term safety and personalization become paramount. The risk of off-target effects from gene editors like CRISPR/Cas9 is a primary concern, necessitating comprehensive off-target analysis in pre-clinical models [3] [87]. Additionally, the high genetic variability of HIV and the heterogeneity of the latent reservoir among individuals demand personalized treatment frameworks. This may involve pre-screening a patient's viral tropism and HLA type to tailor the editing strategy—for example, deciding whether a multi-target approach against CXCR4 is necessary or which bNAbs would be most effective against the patient's specific viral quasispecies [3]. The ultimate goal is to develop a one-time autologous therapy that is both highly effective and broadly accessible, moving beyond the serendipitous cures of the past towards a reliable and scalable medical solution for HIV.
The C-C chemokine receptor type 5 (CCR5) serves as a pivotal co-receptor for human immunodeficiency virus (HIV) entry into CD4+ T-cells, making it a cornerstone target for curative strategies. The discovery that a natural 32-base pair deletion in the CCR5 gene (CCR5-Δ32) confers profound resistance to R5-tropic HIV-1 infection in homozygous individuals provided a genetic blueprint for novel therapies. This was unequivocally demonstrated by the cases of the "Berlin" and "London" patients, who achieved sustained viral remission following hematopoietic stem cell transplantation from CCR5-Δ32 homozygous donors [3] [89]. However, the global distribution of the CCR5-Δ32 allele is highly heterogeneous, with a frequency of approximately 10% in Northern European populations but being virtually absent in African, Asian, and Native American populations [90] [11]. This geographic disparity, combined with the challenges of finding HLA-matched donors, renders allogeneic transplantation an unviable widespread solution, thereby spurring the development of gene editing technologies to mimic this protective mutation in patient-derived cells.
Current antiretroviral therapy (ART), while effective at suppressing viral replication, fails to eradicate latent viral reservoirs and necessitates lifelong adherence, posing challenges of cumulative drug toxicity, cost, and potential resistance development [3] [89]. Gene editing approaches, particularly those utilizing CRISPR/Cas9, aim to overcome these limitations by creating a durable, intrinsic resistance to HIV infection. The primary challenge now lies in transitioning these promising proof-of-concept therapies from boutique laboratory procedures to scalable, cost-effective, and globally accessible treatments. This whitepaper examines the economic and manufacturing hurdles inherent in this transition and outlines the technological innovations and strategic frameworks necessary to overcome them.
Technical Foundations: Gene Editing Technologies and Workflows
Key Gene Editing Platforms for CCR5 Disruption
Several nuclease platforms have been developed to precisely target and disrupt the CCR5 gene. Each technology offers distinct advantages and limitations in terms of specificity, efficiency, and manufacturability.
Table 1: Comparison of Major Gene Editing Platforms for CCR5 Targeting
| Technology | Mechanism of Action | Advantages | Limitations and Challenges | Clinical Stage |
|---|---|---|---|---|
| Zinc Finger Nucleases (ZFNs) | Custom-designed zinc finger proteins fused to FokI nuclease dimerize to induce DNA cleavage. | Early clinical data on safety and efficacy; first CCR5 editor in clinical trials. | Complex design; higher risk of off-target effects; potential immunogenicity. | Clinical trials (e.g., SB-728-T) [3] |
| TALENs | Transcription activator-like effector (TALE) proteins fused to FokI nuclease for DNA cleavage. | More modular design and improved specificity over ZFNs. | Technically demanding construction; large size complicates viral vector delivery. | Preclinical & GMP production established [3] [91] |
| CRISPR/Cas9 | A single guide RNA (sgRNA) directs the Cas9 nuclease to specific genomic loci for cleavage. | Simple design, high efficiency, allows for multiplexed editing. | Off-target effects; PAM sequence dependency; potential for immune responses to prolonged Cas9 expression. | Early-phase clinical trials (e.g., NCT03164135) [3] [53] |
| Base Editors | Fusion of Cas proteins with deaminases enables precise single-nucleotide changes without double-strand breaks. | Avoids risks associated with double-strand breaks (indels, translocations). | Potential for off-target DNA/RNA editing; limited editing window. | Preclinical stage [3] |
Core Experimental Protocols and Workflows
The manufacturing of CCR5-edited cell therapies follows a multi-stage, closed workflow. The protocol below, synthesizing methods from recent studies, outlines a representative process for creating CCR5-negative CD4+ T-cells or Hematopoietic Stem and Progenitor Cells (HSPCs).
Objective: To manufacture clinically relevant doses of CCR5-edited CD4+ T-cells in a GMP-compatible, automated system. Key Reagents and Equipment: CliniMACS Prodigy system (Miltenyi Biotec), GMP-grade CCR5-targeting nuclease mRNA (e.g., TALEN or CRISPR-Cas9), electroporation reagents, cell culture media, and cytokine cocktails [91].
Step-by-Step Protocol:
- Cell Sourcing and Mobilization: CD34+ HSPCs are collected from patients via apheresis after mobilization with Granulocyte Colony-Stimulating Factor (G-CSF) and plerixafor. This combination has been shown to yield high numbers of CD34+ cells and, critically, a significantly higher fraction of long-term engrafting CD90+CD45RA− HSCs compared to bone marrow aspiration [92].
- Cell Selection and Culture: CD4+ T-cells or CD34+ HSPCs are isolated from the leukapheresis product using clinical-grade immunomagnetic selection (e.g., CliniMACS CD4 or CD34 reagent). Cells are then activated and expanded ex vivo using GMP-grade cytokines (e.g., IL-2 for T-cells; SCF, TPO, FLT3-L for HSPCs) [91].
- Gene Editing via Electroporation: Cells are transfected with CRISPR-Cas9 Ribonucleoprotein (RNP) complexes or TALEN mRNA via electroporation. A study using MT4CCR5 cells demonstrated that a RNP complex of 10 µg Cas9 protein and 4 µg of each sgRNA achieved a 97.89% reduction in CCR5 expression [53]. This non-viral method minimizes the risk of genomic integration associated with viral vectors.
- Expansion and Harvesting: Edited cells are cultured in a closed, automated bioreactor system like the CliniMACS Prodigy for large-scale expansion. One automated process reliably produced >1.5 × 10^9 cells with >60% CCR5 editing within 12 days, with about 40% of cells exhibiting biallelic editing [91].
- Product Formulation and Infusion: The final cell product is washed, concentrated, and formulated in an infusion bag. Patients undergo a conditioning regimen (e.g., busulfan) before the edited cells are administered intravenously [92].
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for CCR5 Gene Editing Research
| Reagent / Material | Function in Experimental Protocol | Example Usage |
|---|---|---|
| CCR5-Targeting Nucleases (ZFNs, TALENs, CRISPR-Cas9) | Mediates site-specific DNA cleavage within the CCR5 gene to disrupt its function. | CCR5-Uco-hetTALEN mRNA or CRISPR sgRNAs targeting exon 1 of CCR5 are delivered via electroporation [53] [91]. |
| GMP-Grade mRNA / RNP | In vitro transcribed mRNA or pre-complexed Ribonucleoprotein (RNP) for nuclease delivery. | Enables transient nuclease expression, reducing off-target risks compared to viral DNA delivery [91]. |
| Mobilization Agents (G-CSF, Plerixafor) | Stimulates mobilization of hematopoietic stem cells from bone marrow to peripheral blood for collection. | Administration of G-CSF (10 μg/kg/day for 5 days) with a single dose of Plerixafor (1 mg/kg) on day 4 [92]. |
| CliniMACS Prodigy System | Integrated, closed-system automated cell processing platform. | Used for cell separation, culture, electroporation, and final formulation in a GMP-compliant manner [91]. |
| Conditioning Regimens (Busulfan, Melphalan) | Creates "space" in the bone marrow for the engraftment of newly infused edited cells. | Busulfan conditioning demonstrates significant lymphocyte sparing, preserving adaptive immunity post-transplant [92]. |
| Droplet Digital PCR (ddPCR) | Ultra-sensitive quantification of gene editing efficiency and off-target effects. | Used with specific primers/probes (e.g., CCR5ref, CCR5mut) to precisely measure indel frequencies at the on-target CCR5 locus [91]. |
Manufacturing and Scalability Challenges
Scaling the production of CCR5-edited therapies to meet global demand confronts significant biological and logistical bottlenecks that directly impact both efficacy and cost.
Process-Related Hurdles
A primary challenge is the high variability of starting biological material. Donor-to-donor differences in cells lead to unpredictable performance during manufacturing and inconsistent final product quality [93]. Furthermore, maintaining the therapeutic potency of cells during expansion is difficult. As one expert notes, "While we can grow large numbers of CAR-T cells, maintaining their stemness and preventing exhaustion during manufacturing remains difficult, directly impacting patient outcomes" [93]. This is particularly relevant for creating persistent, functional CCR5-negative immune cells.
The dominant model for autologous therapies has been centralized manufacturing, which creates a complex, patient-specific supply chain. This model introduces severe challenges in cold-chain maintenance, strict time constraints, and the critical need for end-to-end traceability [93]. The "vein-to-vein" process is further strained by a shortage of specialized professionals and QC testing constraints, which can delay product release [93].
The Economic Burden of Manufacturing
The cost of goods sold (COGS) for autologous cell therapies remains prohibitively high. Legacy manufacturing processes are "complex, resource-intensive, and difficult to scale," creating a bottleneck that is the "leading driver of high therapeutic costs" [93]. These costs are driven by expensive raw materials, labor-intensive open processes, and the need for dedicated cleanroom facilities. The high price tag of existing cell and gene therapies (ranging from hundreds of thousands to millions of dollars per treatment) creates immense pressure on healthcare systems and struggles with "global mechanisms for pricing and reimbursement" [93]. Achieving commercial viability requires a fundamental re-evaluation of these processes to lower costs without compromising quality.
Strategies for Enhancing Global Accessibility
Overcoming the challenges of scalability and cost requires a multi-pronged approach focusing on technological innovation, novel supply chain models, and strategic financial planning.
Technological and Process Innovations
- Automation and Closed Systems: Implementing automated, closed-system platforms like the CliniMACS Prodigy reduces manual handling, decreases contamination risk, improves process consistency, and lowers labor costs [91]. As noted by industry experts, "Adopting new and emerging technologies will be important and the ability to automate complex processes will be critical to drive down costs and meet the demand of larger patient populations" [93].
- Novel Delivery Systems: Innovations in drug delivery, such as hydrogel encapsulation, have the potential to reduce manufacturing complexity and simplify logistics by eliminating the need for cryopreservation [93].
- In Vivo Gene Editing: This disruptive approach involves directly delivering the gene editing machinery to a patient's cells in vivo, bypassing the need for complex ex vivo manufacturing altogether. While this technology is still emerging, it "holds great promise for patients" by potentially dramatically simplifying the therapeutic pipeline [93].
Novel Supply Chain and Manufacturing Models
To address the limitations of centralized production, the industry is exploring fit-for-purpose models.
- Decentralized and Point-of-Care Manufacturing: Establishing regional or hospital-based manufacturing centers can drastically reduce transport times and logistics costs, improving patient access in underserved regions [93]. However, this requires overcoming hurdles in site accreditation and standardizing processes across multiple locations.
- Supply Chain Digitization: Advanced digital platforms are crucial for managing the patient-specific supply chain, providing end-to-end visibility, and maintaining an unbreakable chain of identity and custody from collection to infusion [93].
Economic and Accessibility Considerations
- Addressing High Costs: Beyond manufacturing innovations, simplifying therapeutic processes wherever possible is key to reducing costs. Additionally, developing innovative payment models, such as outcome-based agreements and installment plans, can help align the high upfront cost with the long-term therapeutic benefit and potential reduction in lifetime ART expenses [93].
- Ensuring Global Access: Making these therapies accessible worldwide requires tackling infrastructure disparities and a lack of harmonized regulations. "Bridging this gap will require innovative delivery models, such as decentralized manufacturing and point-of-care solutions," especially for underserved populations with limited access to large clinical centers [93]. Engaging patient advocacy groups and working towards regulatory standardization across countries are also critical steps.
The path to a functional cure for HIV via CCR5 gene editing is scientifically clearer than ever, but its realization on a global scale depends on solving formidable manufacturing and economic challenges. The success of this endeavor hinges on the sector's ability to replace legacy processes with automated, scalable, and robust manufacturing platforms; to re-imagine the supply chain through decentralized and point-of-care models; and to collaborate across industry, academia, and regulatory bodies to create sustainable pricing and reimbursement frameworks.
Future efforts must focus on integrating multi-target editing strategies (e.g., simultaneously targeting CCR5 and CXCR4 to prevent tropism switching) with immunotherapies to enhance efficacy [3] [53]. Concurrently, continuous optimization of conditioning regimens, like the lymphocyte-sparing busulfan protocol [92], will improve safety and treatment outcomes. By addressing the dual imperatives of scientific innovation and pragmatic scalability, the research community can transform CCR5 gene editing from a transformative concept into a practical and accessible solution for millions of people living with HIV worldwide.
Evaluating Efficacy and Safety: Weighing CCR5-Targeted Approaches Against Standards of Care
The C-C chemokine receptor type 5 (CCR5) serves as a major co-receptor for human immunodeficiency virus type 1 (HIV-1) entry into host cells, playing a pivotal role in viral pathogenesis [94]. The discovery that a 32-base-pair deletion (Δ32) in the CCR5 gene can confer resistance to HIV-1 infection represented a breakthrough in understanding host-genetic factors in viral susceptibility [38]. This meta-analysis systematically quantifies the protective effect of the CCR5-Δ32 polymorphism against HIV-1 infection, providing researchers and drug development professionals with a comprehensive evidence synthesis. The findings are framed within the broader context of CCR5 research, which has evolved from fundamental genetic discovery to innovative therapeutic strategies including gene editing and CCR5 blockade [3] [87].
Quantitative Synthesis of Evidence
A comprehensive meta-analysis incorporating 24 case-control studies with 4,786 HIV-1 infected patients and 6,283 controls revealed distinct susceptibility patterns based on genotype [12]. The analysis demonstrated that the protective effect varies significantly according to zygosity status, with homozygous individuals showing the most pronounced resistance to infection.
Table 1: Pooled Odds Ratios for HIV-1 Infection Susceptibility by CCR5-Δ32 Genotype
| Genotype | Comparison Group | Odds Ratio (OR) | 95% Credible Interval | Protective Effect |
|---|---|---|---|---|
| Δ32 Heterozygotes | Wild-type homozygous | 1.16 | 1.02 - 1.32 | Increased susceptibility |
| Δ32 Homozygous | Wild-type homozygous | 0.25 | 0.09 - 0.68 | Significant protection |
| Δ32 Allele Carriers | Exposed uninfected | 0.71 | 0.54 - 0.94 | Significant protection |
Ethnic Distribution and Population Stratification
The frequency of the CCR5-Δ32 polymorphism demonstrates substantial geographic and ethnic variation, with highest prevalence observed in Caucasian populations of Northern European descent [12] [38]. This distribution pattern has important implications for the global applicability of research findings and therapeutic approaches targeting CCR5.
Recent research from Angola found a complete absence of the CCR5-Δ32 allele among 272 participants, underscoring the mutation's rarity in African populations [29]. In contrast, studies in European-derived populations report allele frequencies of approximately 10%, with homozygosity occurring in about 1% of these populations [38].
Molecular Mechanisms of Protection
CCR5 Structure and Viral Entry
CCR5 is a G protein-coupled receptor characterized by seven transmembrane helices, an extracellular N-terminus, three extracellular loops (ECLs), and an intracellular C-terminus [95]. Key structural elements for HIV-1 interaction include the N-terminus and second extracellular loop, which facilitate binding with the viral envelope glycoprotein gp120 [95].
The 32-base-pair deletion results in a truncated, non-functional receptor that fails to reach the cell surface, thereby preventing HIV-1 from utilizing it as an entry co-receptor [38] [94]. This molecular mechanism explains the profound resistance observed in homozygous individuals, particularly against R5-tropic HIV-1 strains that dominate during early and chronic infection phases [3].
Diagram 1: Molecular mechanism of CCR5-Δ32 mediated HIV-1 resistance (Title: CCR5-Δ32 HIV Resistance Mechanism)
Coreceptor Switching and Limitations
While CCR5-Δ32 homozygosity provides robust protection against R5-tropic HIV-1 strains, this protection is not absolute. Case reports have documented HIV-1 infection in CCR5-Δ32 homozygous individuals, typically involving viral variants that utilize alternative coreceptors, particularly CXCR4 [3] [87]. This phenomenon, known as coreceptor switching, represents a significant limitation of strategies focused exclusively on CCR5 blockade.
Recent research has revealed that X4-tropic variants are predominantly observed in late-stage infection and are associated with different immune activation patterns compared to R5-tropic viruses [96]. Studies in Mexico City found that 18.4% of treatment-naïve patients harbored mixed R5/X4 viral populations, highlighting the clinical relevance of this viral diversity [96].
Methodological Approaches
Genotyping Techniques
The primary studies included in the meta-analysis employed varied but well-established methodologies for CCR5-Δ32 detection. Most utilized polymerase chain reaction (PCR)-based approaches, with some implementing restriction fragment length polymorphism (PCR-RFLP) analysis for enhanced specificity [12].
Table 2: Essential Research Reagents for CCR5-Δ32 Genotyping
| Research Reagent | Function | Specification Notes |
|---|---|---|
| DNA Extraction Kit | Genomic DNA isolation | QIAamp DNA Mini Kit or equivalent |
| CCR5-specific Primers | Amplification of target region | Flanking Δ32 deletion site |
| PCR Master Mix | DNA amplification | Contains Taq polymerase, dNTPs, buffer |
| Agarose Gel | Electrophoretic separation | 2% concentration for clear resolution |
| DNA Size Marker | Fragment size determination | Including 262bp wild-type reference |
Advanced Sequencing Technologies
Next-generation sequencing (NGS) platforms have enabled more comprehensive analysis of CCR5 and related genes. A recent Turkish study utilized targeted NGS (tNGS) to examine CCR5, CXCR4, and IFNAR1 gene variations, achieving 100% coverage of targeted bases with >100× sequencing depth [97]. This approach facilitates the identification of both common polymorphisms and rare variants that may influence HIV-1 disease progression.
The typical workflow for such analyses includes: (1) genomic DNA extraction from whole blood, (2) amplification of target regions via long-range PCR, (3) library preparation using tagmentation-based methods, (4) sequencing on platforms such as Illumina NovaSeq 6000, and (5) bioinformatic analysis using standardized pipelines like GATK Best Practices [97].
Diagram 2: Next-generation sequencing workflow for CCR5 genotyping (Title: NGS CCR5 Genotyping Workflow)
Therapeutic Applications and Research Translation
CCR5-Targeted Therapies
The protective effect of CCR5-Δ32 has inspired multiple therapeutic approaches aimed at mimicking this natural resistance mechanism. Maraviroc, a small-molecule CCR5 antagonist, is currently the only licensed CCR5 inhibitor approved for treatment-naïve and treatment-experienced patients infected with CCR5-tropic HIV-1 [95]. Additional candidates, including Cenicriviroc (CVC), are in advanced clinical development [95].
Gene editing technologies represent a more permanent therapeutic strategy. CRISPR-Cas9, zinc finger nucleases (ZFNs), and transcription activator-like effector nucleases (TALENs) have all demonstrated efficacy in disrupting the CCR5 gene ex vivo [3] [87]. Clinical trials investigating ZFN-modified CD4+ T cells (NCT03666871) and CRISPR-Cas9-edited hematopoietic stem cells (NCT03164135) have shown promising results in establishing HIV-1 resistance [3].
Curative Strategies and Combination Approaches
Allogeneic transplantation of CCR5-null hematopoietic stem and progenitor cells (HSPCs) remains the only documented cure for HIV-1 infection, as demonstrated in the "Berlin" and "London" patients [3] [87]. However, this approach is limited by donor availability and transplantation-associated risks.
Innovative strategies now aim to combine CCR5 knockout with other antiviral mechanisms in autologous HSPCs. A recent breakthrough demonstrated multilayered HIV-1 resistance through CCR5 knockout coupled with B cell secretion of broadly neutralizing antibodies (bNAbs) such as 10-1074, PGDM1400, and Ibalizumab [87]. This approach addresses the limitation of CXCR4-tropic viral escape while potentially providing both cell-intrinsic and cell-extrinsic protection.
This meta-analysis quantitatively establishes that CCR5-Δ32 homozygosity confers significant protection against HIV-1 infection, with an odds ratio of 0.25 compared to wild-type homozygous individuals. The mutation's effect exhibits dose-dependency, with heterozygosity showing modest influence on susceptibility. These genetic insights have catalyzed the development of CCR5-targeted therapeutic strategies that recapitulate the protective effect in infected individuals.
Future research directions should focus on multi-target strategies addressing both CCR5 and CXCR4 coreceptors, personalized approaches accounting for viral tropism and host genetics, and safety-optimized gene editing platforms with enhanced global accessibility [3]. The continued investigation of CCR5 biology and the Δ32 mutation remains crucial for advancing both functional cure strategies and preventive interventions for HIV-1.
This whitepaper synthesizes clinical and preclinical outcomes of Zinc Finger Nuclease (ZFN) and CRISPR/Cas9 gene editing technologies targeting the CCR5 co-receptor for HIV therapy. Framed within the broader context of CCR5 biology and the protective CCR5-Δ32 mutation, we present comprehensive efficacy and safety data from published trials and studies. The data reveal that both platforms demonstrate successful engraftment of edited cells, reduced viral replication, and encouraging safety profiles, though with distinct efficiency and risk characteristics. Critical challenges including viral tropism switching, variable engraftment efficiency, and off-target effects are examined. This analysis provides researchers and drug development professionals with a technical foundation for selecting and optimizing gene editing approaches for HIV functional cure strategies.
The C-C chemokine receptor type 5 (CCR5) serves as a crucial co-receptor for human immunodeficiency virus (HIV) entry into CD4+ T-cells and macrophages. The discovery that individuals carrying a homozygous 32-base pair deletion in the CCR5 gene (CCR5-Δ32/Δ32) exhibit natural resistance to R5-tropic HIV-1 strains provided the fundamental rationale for CCR5-targeted gene therapies [21] [98]. This mutation results in a truncated protein that fails to localize to the cell surface, thereby preventing viral docking and entry [98]. The clinical proof-of-concept was established through the "Berlin," "London," and additional patients who achieved sustained HIV remission following hematopoietic stem cell transplantation from CCR5-Δ32 homozygous donors [21] [53].
Despite this validation, the scarcity of matched CCR5-Δ32 donors (approximately 1% in Caucasian populations) necessitates alternative approaches to generate HIV-resistant cells [53]. Gene editing technologies—particularly ZFNs and CRISPR/Cas9—have emerged as promising strategies to recapitulate this protective phenotype in patient-derived cells. This whitepaper examines the clinical and preclinical outcomes of these approaches, focusing on efficacy, safety, and technical implementation.
Clinical and Preclinical Outcomes of ZFN and CRISPR/Cas9 Editing
Quantitative Efficacy and Safety Data from Key Studies
Table 1: Clinical and Preclinical Outcomes of CCR5-Targeted Gene Editing
| Study Reference | Technology | Editing Efficiency | HIV Challenge Results | Engraftment/Safety Observations |
|---|---|---|---|---|
| NCT03164135 (Clinical Trial) [21] [53] | CRISPR/Cas9 (HSPCs) | Successful CCR5 ablation | Persistent CCR5 ablation in peripheral blood for >19 months | No gene-editing-related adverse events detected; stable engraftment |
| ZFN SCD Trial (BIVV003) [99] | ZFN (HSPCs) | 75.3% (healthy donors), 64.2% (SCD donors) allele modification | N/A (for SCD) | Well-tolerated; no severe vaso-occlusive crises; long-term multilineage engraftment in vivo |
| Preclinical (CEM Cells) [100] | CRISPR/Cas9 (Lentiviral) | 87.9% indel frequency; 4.3% CCR5+ cells vs. 81.7% in control | >100-fold reduction in p24 antigen vs. control | N/A (in vitro study) |
| Preclinical (Macrophages) [100] | CRISPR/Cas9 (CD34+ HSPCs) | Significant CCR5 disruption | 10-25 fold reduction in p24 antigen levels | Successful differentiation into HIV-resistant macrophages |
| Preclinical (MT4CCR5 Cells) [53] | CRISPR/Cas9 (RNP) | Dose-dependent: 10.43% CCR5+ (low dose) vs. 1.91% CCR5+ (high dose) | Protection against R5-tropic HIV | 77.5-98.4% cell viability post-nucleofection |
Table 2: Technology Comparison for CCR5-Targeted HIV Therapy
| Technology | Mechanism of Action | Advantages | Limitations and Challenges | Clinical Stage |
|---|---|---|---|---|
| ZFN [21] [99] | Custom zinc finger proteins fuse to FokI nuclease for targeted cleavage | Early clinical trial entry; accumulated safety/efficacy data | Complex design; higher off-target risk; potential immunogenicity | Clinical trials (Phase 1/2 for SCD) |
| CRISPR/Cas9 [21] [53] [100] | sgRNA guides Cas9 nuclease to specific genomic loci | Easy design; high efficiency; multiplex editing capability | Off-target effects; PAM sequence dependency; potential immune responses | Early-phase clinical trials |
Analysis of Efficacy Endpoints
ZFN Clinical Outcomes: The BIVV003 trial for sickle cell disease (relevant for HIV due to shared HSPC editing platforms) demonstrated high allelic modification rates (75.3% in healthy donors, 64.2% in SCD donors) with preserved cell viability and function [99]. Edited HSPCs maintained long-term multilineage engraftment potential in NSG mouse models, with stable editing levels observed 19 weeks post-transplantation across lymphoid and myeloid lineages [99].
CRISPR/Cas9 Clinical and Preclinical Outcomes: The first clinical application of CRISPR/Cas9 for CCR5 editing (NCT03164135) in HIV-infected patients with acute lymphoblastic leukemia demonstrated successful engraftment with persistent CCR5 ablation in peripheral blood for over 19 months without gene-editing-related adverse events [21] [53]. Preclinical studies show dose-dependent CCR5 knockout efficiency, with high-dose RNP complexes reducing CCR5 expression to 1.91% compared to 99.80% in controls [53]. HIV challenge experiments consistently show 10-100 fold reductions in viral replication markers across cell lines and primary cells [53] [100].
Experimental Protocols and Methodologies
ZFN-Mediated Gene Editing Protocol
The BIVV003 clinical trial utilized the following methodology for ZFN-mediated BCL11A enhancer editing in HSPCs [99]:
- Stem Cell Mobilization and Collection: HSPCs were mobilized using plerixafor alone (for SCD patients) or plerixafor plus granulocyte colony-stimulating factor (G-CSF) for healthy donors.
- ZFN Delivery: Isolated CD34+ HSPCs were transfected with mRNA encoding ZFNs targeting the GATAA motif in the BCL11A erythroid-specific enhancer.
- Editing Verification: Surveyor Nuclease Assay and next-generation sequencing (NGS) confirmed on-target indel frequencies. For single-cell analysis, edited HSPCs were sorted into single cells and genotyped after erythroid differentiation.
- Product Formulation and Transplantation: Edited cells were formulated as BIVV003 and cryopreserved. Patients received myeloablative busulfan conditioning before autologous transplantation of the gene-edited product.
- Efficacy Assessment: Engraftment success, HbF levels, and clinical outcomes (e.g., vaso-occlusive crises) were monitored post-transplantation.
CRISPR/Cas9-Mediated CCR5 Knockout Protocol
The methodology for CRISPR/Cas9-mediated CCR5 disruption in MT4CCR5 cells exemplifies a clinically relevant approach [53]:
- RNP Complex Formation: Ribonucleoprotein (RNP) complexes were assembled by combining purified Cas9 protein with synthetic single-guide RNAs (sgRNAs) targeting the first exon of CCR5, corresponding to the Δ32 mutation site.
- Cell Nucleofection: MT4CCR5 cells were nucleofected with RNP complexes using two dosing regimens:
- Low dose: 6μg Cas9 + 2μg of each sgRNA (total 4μg sgRNAs)
- High dose: 10μg Cas9 + 4μg of each sgRNA (total 8μg sgRNAs)
- Efficiency Assessment:
- Cleavage efficiency: T7 Endonuclease I (T7E1) assay at 3 days post-nucleofection.
- Protein expression: Flow cytometry and Western blot analysis of CCR5 surface expression.
- Cell viability: 7-AAD staining and flow cytometry.
- Functional Validation: Edited cells were challenged with R5-tropic HIV-1, with viral replication monitored via p24 antigen ELISA.
ZFN Clinical Workflow
Advanced Technical Considerations
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for CCR5 Gene Editing Studies
| Reagent/Category | Specific Examples | Function and Application |
|---|---|---|
| Gene Editing Platforms | ZFN mRNA, CRISPR/Cas9 RNP | Core editing machinery for targeted DNA cleavage |
| Delivery Systems | Electroporation, Lentiviral Vectors, Virus-like Particles (VLPs) | Facilitate intracellular delivery of editing components |
| Cell Sources | CD34+ HSPCs, Primary T-cells, Cell lines (e.g., MT4CCR5, CEM.NKR-CCR5) | Targets for genetic modification and functional assays |
| Efficiency Validation | T7E1 Assay, Next-generation Sequencing, Flow Cytometry | Quantify editing efficiency and protein disruption |
| Functional Assays | p24 ELISA, Viral Challenge (R5/X4-tropic HIV) | Assess antiviral efficacy of edited cells |
| In Vivo Models | NSG Mice, Humanized Mouse Models | Evaluate engraftment potential and long-term safety |
Addressing Technical Challenges
Overcoming Viral Tropism Switching: A significant limitation of CCR5-only editing is the potential emergence of CXCR4-using (X4-tropic) HIV variants [21] [100]. Multiplexed editing strategies simultaneously targeting CCR5, CXCR4, and viral LTR regions show superior efficacy in establishing comprehensive viral blockade [21]. A preclinical study demonstrated that dual CCR5/CXCR4 disruption conferred resistance to R5-tropic, X4-tropic, and dual-tropic HIV strains [100]. However, CXCR4 editing may impair bone marrow engraftment of CD4+ T cells, necessitating careful evaluation of this approach [100].
DNA Repair Pathway Management: Editing outcomes are heavily influenced by cellular DNA repair mechanisms, which differ significantly between cell types [101]. Non-homologous end joining (NHEJ) dominates in postmitotic cells, while microhomology-mediated end joining (MMEJ) is more prevalent in dividing cells [101]. In neurons, indel accumulation continues for up to two weeks post-editing, contrasting with the rapid resolution (within days) observed in dividing cells [101]. Chemical or genetic manipulation of DNA repair pathways can direct outcomes toward desired edits.
CCR5 in HIV Entry Pathway
Clinical outcomes for ZFN and CRISPR/Cas9 platforms demonstrate measurable progress toward CCR5-targeted HIV therapy. Both technologies show promising efficacy in disrupting CCR5 expression and conferring HIV resistance with acceptable safety profiles in early trials. The choice between platforms involves trade-offs: ZFNs benefit from longer clinical track records, while CRISPR/Cas9 offers simpler design and multiplexing capabilities.
Future HIV cure strategies will likely combine gene editing with complementary approaches. These include:
- Multiplexed gene editing targeting CCR5, CXCR4, and HIV proviral DNA [21]
- Combination with immunotherapy such as CAR-T cells or immune checkpoint inhibitors to enhance viral clearance [21] [3]
- Advanced delivery systems including lipid nanoparticles and VLPs to improve editing efficiency and specificity [64] [101]
- Personalized approaches accounting for viral tropism, host genetics, and latent reservoir characteristics [21]
As these technologies evolve, ongoing attention to long-term safety, off-target effects, and global accessibility will be essential for translating gene editing into widely available HIV therapeutics.
The pursuit of a functional cure for Human Immunodeficiency Virus (HIV) has catalyzed the development of two divergent therapeutic paradigms: lifelong administration of antiretroviral therapy (ART) and the emerging field of gene-editing-based interventions. ART effectively suppresses viral replication but necessitates lifelong daily adherence, presenting challenges related to cumulative toxicity, cost, and persistent latent reservoirs. In contrast, gene-editing strategies, particularly those targeting the C-C chemokine receptor 5 (CCR5)—the principal co-receptor for HIV entry—aim to confer sustained, treatment-free remission through a single or limited number of interventions. This whitepaper provides a technical comparison of the durability, mechanisms, and clinical progress of these approaches, contextualized within the framework of natural CCR5Δ32 mutation research. It further details experimental protocols for evaluating these therapies, visualizes key biological pathways, and catalogues essential research tools for the field.
The C-C chemokine receptor type 5 (CCR5) is a G-protein-coupled receptor expressed on the surface of immune cells, including CD4+ T-cells and macrophages. Its role as the predominant co-receptor for HIV-1 entry during initial infection and early disease stages is well-established [18] [25]. The natural genetic variant CCR5Δ32, a 32-base-pair deletion in the CCR5 gene, results in a truncated protein that is not expressed on the cell surface [23] [20]. Individuals homozygous for this mutation (Δ32/Δ32) are highly resistant to infection with CCR5-tropic HIV strains, while heterozygotes exhibit reduced cell surface CCR5 and slower disease progression [25] [20]. This natural resistance provided the foundational rationale for targeting CCR5 therapeutically. The clinical validation of this approach came from the cases of the "Berlin" and "London" patients, who were functionally cured of HIV after allogeneic hematopoietic stem cell transplantation (allo-HSCT) from CCR5Δ32/Δ32 donors [3]. Gene-editing technologies aim to recapitulate this protective phenotype in a patient's own cells, offering a potential path to a durable, one-time therapy.
Comparative Analysis of Therapeutic Modalities
Mechanism of Action and Durability
Table 1: Mechanism and Durability of ART vs. Gene-Editing Therapies
| Feature | Lifelong Antiretroviral Therapy (ART) | CCR5-Targeted Gene-Editing Therapies |
|---|---|---|
| Primary Mechanism | Pharmacological inhibition of viral enzymes (e.g., reverse transcriptase, integrase, protease) to suppress viral replication [3]. | Permanent disruption of the CCR5 gene via nucleases (e.g., CRISPR-Cas9, ZFNs) or base editors, preventing HIV co-receptor binding and cellular entry [23] [3]. |
| Therapeutic Durability | Transient; requires daily, lifelong adherence to maintain viral suppression. Treatment interruption leads to rapid viral rebound from latent reservoirs [3]. | Aims to be permanent; a single application can generate a long-lasting population of HIV-resistant cells capable of self-renewal (e.g., edited hematopoietic stem cells) [23] [3]. |
| Impact on Latent Reservoir | None. ART suppresses active replication but does not eliminate latently infected cells, which form a stable reservoir [3]. | Indirectly targets the reservoir by preventing new infections. Can be combined with strategies to directly target integrated provirus (e.g., editing HIV LTR) [3]. |
| Evidence of Durability | Well-established for the duration of treatment. No sustained remission upon cessation without additional interventions [88]. | Demonstrated in humanized mouse models with long-term HIV resistance [23]. Case reports of cured individuals post-CCR5Δ32/Δ32 allo-HSCT show durability over many years [3]. |
Clinical Outcomes and Risks
Table 2: Clinical Profile and Challenges of HIV Therapies
| Aspect | Lifelong Antiretroviral Therapy (ART) | CCR5-Targeted Gene-Editing Therapies |
|---|---|---|
| Efficacy | Highly effective at suppressing plasma viremia to undetectable levels, restoring immune function, and preventing AIDS [3]. | Proof-of-concept established in allo-HSCT settings and early-phase trials. Editing efficiency and the size of the protected cell population are critical determinants of success [23] [3]. |
| Primary Challenges | Lifelong adherence, cumulative drug toxicity, drug-drug interactions, emergence of drug resistance, and financial cost [3]. | Technical challenges include: risk of off-target mutations; mosaicism in edited embryos; and potential for immune rejection of edited cells [23] [102] [3]. |
| Safety Profile | Generally well-characterized with known side-effect profiles (e.g., metabolic, renal, hepatic). | Long-term safety is still under investigation. A key concern is the biphasic role of CCR5; while protective against HIV, its absence may increase susceptibility to other pathogens like West Nile virus and influenza, and impact recovery from stroke [23] [25]. |
Experimental Protocols for Evaluating Gene-Editing Therapies
Protocol 1: CCR5 Ablation in Hematopoietic Stem/Progenitor Cells (HSPCs)
This protocol is foundational for developing a durable, single-treatment therapy aimed at generating a complete HIV-resistant immune system [23] [3].
- Cell Isolation and Culture: Isolate human CD34+ HSPCs from mobilized peripheral blood or cord blood using magnetic-activated cell sorting (MACS) or fluorescence-activated cell sorting (FACS). Culture cells in serum-free medium supplemented with cytokines (SCF, TPO, FLT3-L).
- Delivery of Gene-Editing Machinery:
- CRISPR-Cas9: Electroporation of ribonucleoprotein (RNP) complexes comprising Cas9 nuclease and a synthetic guide RNA (gRNA) targeting exon 1 or 3 of the CCR5 gene. Alternatively, use lentiviral vectors to deliver Cas9 and gRNA expression cassettes [3].
- Zinc Finger Nucleases (ZFNs): Electroporation of plasmid DNA or mRNA encoding ZFNs specific for the CCR5 gene (e.g., SB-728-T clinical trial construct) [3].
- In Vitro Validation:
- Editing Efficiency: Assess by T7 Endonuclease I assay or Tracking of Indels by Decomposition (TIDE) analysis. Quantify CCR5 protein loss via flow cytometry using anti-CCR5 antibodies.
- Functional Assay: Challenge edited cells with CCR5-tropic HIV-1 (e.g., Bal. strain) and measure p24 levels or intracellular HIV RNA to confirm resistance.
- In Vivo Durability Assessment:
- Transplant edited CD34+ cells into immunodeficient NSG mice.
- Monitor long-term engraftment and multi-lineage differentiation in bone marrow and peripheral blood over 4-6 months.
- Re-challenge the humanized mice with HIV and monitor viral load and CD4+ T-cell count. The sustained presence of edited, HIV-resistant cells demonstrates durability [23].
Protocol 2: Assessing Multi-Target Editing Strategies
To combat viral escape via tropism switching, this protocol involves simultaneous editing of multiple host and viral targets [3].
- Multiplexed Guide RNA Design: Design and clone gRNAs targeting CCR5, CXCR4 (the alternative HIV co-receptor), and regulatory regions of the HIV provirus (e.g., Long Terminal Repeat - LTR) into a single CRISPR vector or as multiple RNP complexes.
- Co-delivery and Editing: Electroporate primary CD4+ T cells or HSPCs with the multiplexed editing reagents.
- Comprehensive Analysis:
- Efficiency and Specificity: Use deep sequencing to quantify editing rates at all on-target loci and computationally predicted off-target sites.
- Viral Challenge: Infect cells with a panel of R5-tropic, X4-tropic, and dual-tropic HIV-1 strains. The combination of CCR5 and CXCR4 editing should confer broad resistance against all tropisms.
- Reservoir Activation: Treat edited, laterally infected cell lines with latency-reversing agents (e.g., PMA/ionomycin). Cells with disrupted HIV LTR should show impaired viral reactivation and particle production [3].
Visualization of Key Concepts
HIV Entry and CCR5Δ32 Mechanism
Gene-Editing Experimental Workflow
The Scientist's Toolkit: Key Research Reagents
Table 3: Essential Reagents for CCR5 and HIV Gene-Editing Research
| Reagent / Solution | Function / Application | Technical Notes |
|---|---|---|
| CD34+ MicroBead Kit (e.g., Miltenyi Biotec) | Immunomagnetic isolation of human hematopoietic stem/progenitor cells from source material. | Critical for obtaining a pure cell population for HSPC editing and transplantation studies. |
| CRISPR-Cas9 Ribonucleoprotein (RNP) | Direct delivery of pre-complexed Cas9 protein and synthetic gRNA for high-efficiency, transient editing. | Reduces off-target effects and immune responses compared to plasmid-based delivery [3]. |
| Anti-CCR5 Antibody (clone 2D7) | Flow cytometric analysis of CCR5 cell surface expression to quantify editing efficiency. | Binds to the second extracellular loop of CCR5, which is critical for HIV gp120 binding [23]. |
| R5-tropic HIV-1 Stock (e.g., Bal., SF162) | Functional validation of CCR5 ablation by challenging edited cells in vitro or in humanized mouse models. | Essential for demonstrating phenotypic resistance post-editing. |
| NSG (NOD-scid-IL2Rγnull) Mice | In vivo model for human immune system reconstitution (HSC engraftment) and HIV challenge studies. | Supports long-term engraftment of edited human cells, allowing for durability assessment [23]. |
| T7 Endonuclease I / TIDE Analysis Software | Detection and quantification of insertion/deletion (indel) mutations at the target genomic locus. | Standard methods for initial, rapid evaluation of gene-editing efficiency. |
The comparative durability of gene-editing therapies and lifelong ART represents a fundamental shift from chronic viral suppression to potential sustained remission or cure. While ART remains a life-saving cornerstone of HIV management, its requirement for lifelong daily adherence underscores the need for more durable solutions. CCR5-targeted gene editing, inspired by the natural CCR5Δ32 mutation, holds immense promise for a one-time, curative intervention by creating a permanently protected immune system.
Significant challenges remain, including optimizing editing efficiency and safety, managing the biphasic nature of CCR5 deletion, and making these therapies globally accessible. Future research will focus on combining CCR5 editing with other strategies, such as CXCR4 disruption, direct "shock and kill" of latent reservoirs, and immunotherapy, to construct an insurmountable barrier to HIV infection and rebound. The ongoing refinement of gene-editing platforms, including the advent of base editing with its reduced risk of double-strand breaks, will be crucial in translating this durable therapeutic paradigm from the laboratory to the clinic.
The CC chemokine receptor 5 (CCR5) serves as the primary co-receptor for human immunodeficiency virus type 1 (HIV-1) entry into host cells, establishing it as a critical determinant of viral transmission and pathogenesis in humans [103] [43]. This biological pathway has been further validated by the discovery of the CCR5Δ32 polymorphism, a 32-base-pair deletion that results in a loss-of-function mutation, providing homozygous individuals with significant resistance to HIV-1 infection [12] [103]. Interestingly, parallel evolution of CCR5-null phenotypes has been documented in natural hosts of simian immunodeficiency virus (SIV), particularly in sooty mangabey monkeys, which experience non-pathogenic SIV infection despite high viral replication levels [104]. This remarkable convergence suggests that similar evolutionary pressures have shaped CCR5 biology across species and highlights the potential for natural host studies to inform therapeutic development.
Natural hosts of SIV, including sooty mangabeys and African green monkeys, exhibit a fundamentally different relationship with their respective immunodeficiency viruses compared to the pathogenic infection observed in humans or macaques [105]. Despite robust viral replication, these animals typically maintain normal CD4+ T-cell counts and avoid the progressive immunodeficiency that characterizes AIDS. Emerging evidence indicates that this non-pathogenic outcome may be linked to fundamental differences in viral entry mechanisms, specifically the utilization of alternative coreceptors beyond CCR5 [105]. This review examines the parallel evolution of CCR5 modulation in natural SIV hosts, analyzes the underlying mechanisms that confer protection against disease pathogenesis, and explores the implications for developing novel therapeutic strategies targeting HIV-1 in humans.
Coreceptor Specificity and Cell Targeting: Contrasting Outcomes in Natural versus Non-Natural Hosts
The Established Dogma of CCR5-Mediated Entry
Traditional understanding of immunodeficiency virus entry has centered on CCR5 as the essential coreceptor for viral fusion with target cells. In non-natural host infections, including HIV-1 infection of humans and SIVmac infection of rhesus macaques, this CCR5-dependent entry pathway directs the virus to critical populations of CD4+ T-cells located in lymphoid tissues, which are essential for maintaining immune homeostasis and gut barrier integrity [105]. The targeting of these strategically important cell populations ultimately results in progressive immune depletion and the development of AIDS. The central role of CCR5 in pathogenic infection is further substantiated by the observed resistance to HIV-1 infection in individuals homozygous for the CCR5Δ32 allele, demonstrating the critical nature of this entry pathway for establishing infection in humans [12] [103].
Alternative Coreceptor Usage in Natural Hosts
Contrary to the established paradigm, natural host infections are characterized by a more flexible approach to viral entry. While natural hosts express exceedingly low levels of CCR5, they maintain high levels of virus replication, suggesting the involvement of alternative coreceptors [105]. Research indicates that in multiple natural host species, CCR5 is dispensable for SIV infection both ex vivo and in vivo. Instead, emerging data point to alternative coreceptors, particularly CXCR6, playing a central role in infection and cell targeting [105]. This fundamental difference in viral entry mechanism has profound implications for disease outcome, as it appears to direct the virus to distinct populations of CD4+ T-cells that are more dispensable for immune homeostasis, particularly extralymphoid and more differentiated CD4+ T-cells [105].
Table 1: Coreceptor Usage and Infection Outcomes in Natural versus Non-Natural Hosts
| Characteristic | Natural Hosts (e.g., Sooty Mangabeys) | Non-Natural Hosts (e.g., Humans, Macaques) |
|---|---|---|
| Primary Coreceptor | Alternative coreceptors (e.g., CXCR6); CCR5 dispensable | CCR5-exclusive entry |
| CCR5 Expression | Exceedingly low levels | High levels on target cells |
| Cell Populations Targeted | Extralymphoid, differentiated CD4+ T-cells | Lymphoid tissue CD4+ T-cells critical for immune homeostasis |
| Disease Outcome | Non-pathogenic despite high viremia | Progressive immunodeficiency (AIDS) |
| Genetic Resistance | CCR5-null alleles identified [104] | CCR5Δ32 polymorphism provides resistance [12] [103] |
Implications of Differential Cell Targeting
The shift in coreceptor usage from CCR5 to alternative receptors like CXCR6 in natural hosts represents more than just a viral adaptation—it fundamentally alters the cellular tropism of the virus and its subsequent impact on the immune system. By bypassing CCR5, SIV in natural hosts infects cell populations that are less critical for maintaining overall immune function, thereby preserving lymphoid architecture and preventing the massive CD4+ T-cell depletion characteristic of AIDS [105]. This redirection of viral targeting may also contribute to the reduced immune activation observed in natural hosts, even in the face of persistent high-level viral replication. The preservation of gut barrier integrity in natural hosts, in stark contrast to the profound damage observed in pathogenic HIV-1 and SIVmac infections, further underscores the importance of this differential cell targeting in determining disease outcome.
Parallel Evolution of CCR5-Null Phenotypes: Insights from Natural Hosts and Human Populations
CCR5Δ32 in Human Populations
The CCR5Δ32 allele represents one of the most well-characterized examples of human genetic adaptation conferring resistance to infectious disease. This 32-base-pair deletion within the CCR5 coding region results in a truncated, non-functional protein that fails to reach the cell surface, thereby preventing its utilization by HIV-1 for cellular entry [103]. Meta-analyses of case-control studies have demonstrated that individuals homozygous for the CCR5Δ32 polymorphism show significantly reduced susceptibility to HIV-1 infection, with an odds ratio of 0.25 (95%CI=0.09-0.68) compared to those with wild-type homozygous genotypes [12]. The distribution of this allele displays a distinctive geographic pattern, with the highest frequencies found in European populations (approximately 10% allele frequency, 1% homozygous) and a pronounced north-to-south gradient across Europe [103].
The evolutionary origins of the CCR5Δ32 allele have been the subject of extensive investigation, with several historical selective pressures proposed, including bubonic plague, smallpox, and hemorrhagic fevers [103]. While the specific selective agent remains debated, the high frequency of this allele in certain human populations, despite potential negative consequences such as increased severity to West Nile virus infection, suggests strong historical selective pressure from pathogens that exploited the CCR5 receptor [103].
Parallel CCR5 Mutations in Sooty Mangabeys
Strikingly, a parallel evolutionary phenomenon has been documented in sooty mangabeys, natural hosts of SIV. Research has identified a novel deletion allele of CCR5 in this species, with an allele frequency of approximately 0.04 [104]. Similar to the human CCR5Δ32 variant, the mutant protein in mangabeys is not expressed at the cell surface and consequently does not function as a viral coreceptor. Primary activated lymphocytes from mangabeys heterozygous for the deletion allele expressed significantly less CCR5 on the cell surface [104]. Interestingly, unlike the protective effect observed in humans, SIV seroprevalence and viremia were comparable among CCR5 heterozygotes and wild-type mangabeys, suggesting alternative coreceptor usage allows the virus to bypass this genetic deficiency in natural hosts [104].
Table 2: Epidemiological and Functional Characteristics of CCR5 Null Alleles
| Parameter | Human CCR5Δ32 | Sooty Mangabey CCR5 Deletion |
|---|---|---|
| Genetic Change | 32-bp deletion | Novel deletion allele |
| Protein Expression | Not expressed at cell surface | Not expressed at cell surface |
| Allele Frequency | ~10% in European populations | 0.04 (4%) |
| Homozygous Effect | Highly resistant to HIV-1 infection | Expected to be resistant to CCR5-tropic strains |
| Heterozygous Effect | Reduced cell surface expression; milder AIDS symptoms | Significantly less CCR5 on cell surface |
| Impact on Natural Infection | Protective against HIV-1 acquisition | Comparable SIV seroprevalence and viremia |
| Implied Coreceptor Flexibility | Limited (HIV-1 primarily CCR5-tropic) | High (SIVsmm utilizes alternative coreceptors) |
Evolutionary Implications of Parallel CCR5 Modulation
The independent emergence of CCR5-null alleles in both humans and sooty mangabeys represents a compelling case of parallel evolution and suggests that similar negative selection pressures have acted against CCR5 functionality in these distinct species [104]. This evolutionary convergence indicates that pathogens utilizing CCR5 as an entry portal have likely exerted significant selective pressure throughout mammalian evolution. The different outcomes observed in these species—protection against infection in humans versus maintained infection in mangabeys—highlight the divergent evolutionary trajectories shaped by the availability of alternative viral entry pathways. The mangabey model demonstrates that CCR5-independent infection is not only possible but can result in a non-pathogenic host-virus relationship, providing a paradigm for therapeutic intervention in humans.
Experimental Approaches and Methodologies for Studying Coreceptor Usage
Population Dynamics and Evolutionary Rescue Experiments
Experimental evolution systems using host-virus models have provided valuable insights into the dynamics of parallel evolution and the role of population size changes in shaping evolutionary outcomes. In one such system using algal hosts and viruses, researchers established continuous cultures (chemostats) for approximately 100 host generations, comparing populations evolving alone versus those coevolving with a virus [106]. This approach allowed for direct observation of eco-evolutionary dynamics, where ecology (population size) and evolution (selective sweeps) interact on the same time scale.
The experimental protocol involved:
- Establishing replicate populations: Three coevolved populations (host+virus) and three evolved populations (host alone) were initiated from the same isogenic ancestor.
- Long-term monitoring: Populations were maintained for 90 days with continuous monitoring of host and virus densities.
- Time-shift experiments: Hosts and viruses from different time points were cross-assayed to reconstruct coevolutionary dynamics.
- Wavelet coherence analysis: Statistical method used to test for synchronization of host population dynamics across replicates, revealing significant parallelism in bottlenecks and population expansions.
This methodology revealed that host populations coevolving with viruses underwent significant cycles of population bottlenecks followed by rapid expansions due to evolutionary rescue—the evolution and selective sweeping of new resistant host types [106]. These demographic changes directly influenced patterns of parallel evolution at both phenotypic and genomic levels.
Coreceptor Function and Binding Assays
Understanding the molecular mechanisms of coreceptor function requires detailed binding and functional studies. Key methodological approaches include:
- Binding studies with radiolabeled chemokines: For example, using ¹²⁵I-CCL3 to characterize binding interactions with CCR5 and identify allosteric modulation sites [107].
- Site-directed mutagenesis of CCR5: Systematic alteration of key residues (e.g., Glu-283, Trp-248, Ile-116) to determine their role in receptor function and small molecule interactions [107].
- Cell surface expression quantification: Flow cytometric analysis of CCR5 expression on primary activated lymphocytes from wild-type versus heterozygous animals [104].
- Viral entry assays: Measuring infection efficiency using vectors pseudotyped with envelope proteins from different viral strains in cells with varying coreceptor expression.
These techniques have been instrumental in characterizing the allosteric modulation of CCR5 function and identifying key residues involved in the conformational switches that regulate receptor activation [107].
Genomic Analysis of Evolved Populations
Whole-genome sequencing of evolved host populations has revealed how rapid coevolution shapes genomic architecture. The standard protocol involves:
- Sequencing individual isogenic clones: Multiple clones (e.g., 10) are sequenced from each evolved population and the ancestral population.
- Variant identification: Comparing evolved populations to ancestors to identify derived variants (single-nucleotide polymorphisms and small indels).
- Functional annotation: Classifying variants based on their predicted impact (high, moderate, low, non-coding).
- Parallel evolution assessment: Identifying genomic regions showing repeated evolutionary changes across independent replicate populations.
Application of this approach in host-virus systems has revealed that while phenotypic resistance may evolve in parallel, the underlying genomic changes can show substantial divergence between replicates, with evidence for both parallel duplication of large genomic regions and novel sequence divergence [106].
Data Visualization and Conceptual Framework
Coreceptor Shift in Natural SIV Hosts
The diagram below illustrates the fundamental shift in coreceptor usage that characterizes non-pathogenic SIV infection in natural hosts compared to pathogenic infection in non-natural hosts.
Experimental Workflow for Coreceptor Studies
The following diagram outlines a comprehensive experimental approach for studying coreceptor usage and evolution in natural host systems.
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Key Research Reagents for Studying CCR5 Biology and Coreceptor Function
| Reagent/Category | Specific Examples | Function/Application |
|---|---|---|
| Cell Line Models | Primary lymphocytes, CCR5-transfected cell lines, TZM-bl cells | Viral entry assays, receptor expression studies, infectivity measurements |
| Molecular Biology Tools | Site-directed mutagenesis kits, CCR5-specific antibodies, radiolabeled chemokines (¹²⁵I-CCL3) | Receptor characterization, binding studies, structure-function analysis |
| Chemical Modulators | Maraviroc, ZnBip, ZnTerp, ZnClTerp [107] | Allosteric modulation studies, receptor activation/inhibition assays |
| Genetic Analysis Reagents | CCR5 genotyping assays, whole-genome sequencing kits, RNA-seq reagents | Genetic polymorphism screening, genomic evolution studies, expression profiling |
| Animal Models | Sooty mangabey colonies, African green monkeys, humanized mouse models | Natural host studies, pathogenesis research, therapeutic testing |
| Visualization Tools | Cryo-EM facilities, computational modeling software, flow cytometers | Structural biology, dynamic conformational analysis, receptor quantification |
Therapeutic Implications and Future Directions
Allosteric Modulation of CCR5 Function
The insights gained from natural host studies have profound implications for therapeutic strategies targeting CCR5 in HIV-1 infection. The most successful translation to date has been the development of maraviroc, a non-peptidic, low molecular weight CCR5 ligand that acts as an allosteric modulator [108]. Rather than directly blocking the chemokine binding site, maraviroc stabilizes CCR5 in an inactive conformation that prevents HIV-1 envelope glycoprotein engagement while largely preserving native chemokine signaling functions [108]. Molecular modeling studies have revealed that this allosteric mechanism involves specific interactions with transmembrane domains of CCR5, particularly residues in the major binding pocket that affect the conformational switch of key microswitches like Trp-248 [107].
Recent advances in structural biology, particularly cryo-electron microscopy (cryo-EM), have enabled high-resolution analysis of CCR5 in complex with various ligands, providing a robust foundation for structure-based drug design [43]. These structural insights have facilitated the development of next-generation allosteric modulators with improved efficacy and preservation of CCR5 biological functions. The metal ion chelators ZnBip and ZnTerp complexed with Zn²⁺ represent novel chemotypes that act as CCR5 agonists and positive allosteric modulators of CCL3 binding, offering new avenues for therapeutic intervention [107].
Beyond CCR5: Targeting Alternative Coreceptors
The natural host paradigm demonstrates that shifting viral tropism away from CCR5 represents a viable evolutionary strategy for maintaining immune function despite persistent infection. This suggests that therapeutic approaches aimed at selectively redirecting HIV-1 to utilize alternative coreceptors like CXCR6 might ameliorate disease pathogenesis in humans. While such strategies remain speculative, they represent a radical departure from current therapeutic paradigms focused solely on viral suppression.
Gene editing approaches targeting CCR5 represent another promising therapeutic avenue inspired by natural mutations. The demonstrated protection afforded by the CCR5Δ32 polymorphism has motivated efforts to recreate this phenotype in HIV-1 infected individuals using gene editing technologies like CRISPR-Cas9. While significant technical and ethical challenges remain, this approach represents a direct translation of natural host insights into therapeutic development.
The parallel evolution of CCR5 modulation in natural SIV hosts and human populations provides compelling evidence for convergent evolutionary solutions to immunodeficiency virus infection. The non-pathogenic outcome in natural hosts results from a complex interplay of factors, including alternative coreceptor usage, target cell restriction, and modulated immune activation. By understanding the mechanisms that natural hosts have evolved to coexist with SIV, we can identify novel therapeutic targets and strategies for preventing AIDS in HIV-1 infected humans. The continued investigation of these natural models, combined with advanced structural biology and computational approaches, holds promise for the development of next-generation interventions that mimic the beneficial aspects of natural host adaptations while avoiding their limitations.
The C-C chemokine receptor type 5 (CCR5) represents a paradigm of genetic pleiotropy in human infectious disease. This G protein-coupled receptor, expressed predominantly on T cells, macrophages, and dendritic cells, serves as a critical mediator of immune cell trafficking through its interactions with ligands CCL3, CCL4, and CCL5 [109]. CCR5 gained prominence in virology with the seminal discovery that it functions as the principal coreceptor for macrophage-tropic (R5) human immunodeficiency virus (HIV-1) entry into CD4+ target cells [50]. The subsequent identification of a natural 32-base pair deletion in the CCR5 open reading frame (CCR5Δ32) and its association with profound resistance to HIV-1 infection in homozygous individuals established CCR5 as a therapeutic target of exceptional promise [50].
The paradigm of CCR5 as a dispensable receptor was challenged when research revealed that CCR5 deficiency increases susceptibility to symptomatic West Nile virus (WNV) infection in both mouse models and humans [110] [109]. This discovery unveiled a critical conflict: the same genetic defect that confers HIV-1 resistance simultaneously increases vulnerability to a prevalent neurotropic flavivirus. This risk-benefit dichotomy now represents a central consideration in developing CCR5-targeted therapies, requiring careful analysis for researchers, pharmaceutical developers, and clinical investigators working at the intersection of immunology and infectious disease.
CCR5Δ32: Protective and Pathogenic Effects
Genetic Basis and Population Distribution
The CCR5Δ32 allele contains a 32-base pair deletion within the coding sequence, resulting in a frameshift mutation that generates a truncated protein incapable of trafficking to the cell surface [110]. This null allele follows an autosomal recessive pattern of inheritance, with homozygous individuals (CCR5Δ32/Δ32) exhibiting complete loss of CCR5 function, while heterozygotes show reduced receptor expression [50]. The allelic frequency demonstrates significant geographic and ethnic variation, with highest prevalence in Northern European populations (approximately 10% heterozygous, 1% homozygous), and near absence in African, Asian, and Native American populations [50] [53].
Table 1: CCR5Δ32 Genotype Frequencies and Associated Health Impacts
| Genotype | Receptor Expression | HIV-1 Susceptibility | WNV Disease Risk | Reported Health Status |
|---|---|---|---|---|
| CCR5+/+ (Wild-type) | Normal | Susceptible | Baseline risk | Generally healthy |
| CCR5+/Δ32 (Heterozygous) | Reduced | Delayed progression (2-4 years) | Possibly increased | Generally healthy |
| CCR5Δ32/Δ32 (Homozygous) | Absent | Highly resistant | Significantly increased | Generally healthy, no overt immunodeficiency |
Protective Role Against HIV-1 Infection
The mechanistic basis for HIV-1 resistance in CCR5Δ32 homozygotes stems from the essential role CCR5 plays as a coreceptor during viral entry. HIV-1 envelope glycoproteins sequentially engage CD4 and then CCR5 (or CXCR4) on target cells, triggering fusion and viral entry. In the absence of functional CCR5, R5-tropic viruses—which dominate transmission and early infection—cannot efficiently enter host cells [50]. This protective effect represents one of the strongest genetic associations in infectious disease, with numerous studies confirming that CCR5Δ32 homozygosity confers near-complete resistance to HIV-1 infection despite repeated exposures [110] [50].
The clinical significance of this discovery is underscored by the successful HIV cures achieved in the "Berlin," "London," and "Düsseldorf" patients—HIV-positive individuals with hematological malignancies who received allogeneic hematopoietic stem cell transplantation from CCR5Δ32 homozygous donors [3] [50]. Following transplantation and engraftment of CCR5-deficient immune cells, these patients discontinued antiretroviral therapy and maintained undetectable HIV-1 RNA levels, demonstrating the curative potential of CCR5-targeted approaches [50].
Increased Susceptibility to West Nile Virus Infection
In contrast to its protective effect against HIV-1, CCR5 deficiency confers significantly increased risk for symptomatic WNV infection. Glass et al. first documented this association in 2006, reporting that CCR5Δ32 homozygosity was markedly enriched among symptomatic WNV patients from Arizona and Colorado cohorts compared to healthy controls [110]. Quantitative analysis revealed that CCR5Δ32 homozygotes represented 4.2% and 8.3% of Caucasian WNV patients in the Arizona and Colorado cohorts, respectively, compared to only 1.0% of healthy Caucasian blood donors [110]. The association was particularly strong for fatal outcomes, with CCR5Δ32 homozygotes having an odds ratio of 13.2 for death following WNV infection in the Arizona cohort [110].
Table 2: CCR5Δ32 Homozygosity Frequency in WNV-Infected Cohorts
| Study Cohort | Control Population | Symptomatic WNV Patients | Odds Ratio (95% CI) | P-value |
|---|---|---|---|---|
| Arizona (n=143 Caucasians) | 1.0% (13/1318) | 4.2% (6/143) | 4.4 (1.6-11.8) | 0.0013 |
| Colorado (n=72 Caucasians) | 1.0% (13/1318) | 8.3% (6/72) | 9.1 (3.4-24.8) | <0.0001 |
| Combined (n=215 Caucasians) | 1.0% (13/1318) | 5.6% (12/215) | 5.9 (2.7-12.6) | <0.0001 |
Mouse models have elucidated the immunological mechanism underlying this susceptibility. Following subcutaneous WNV infection, CCR5-deficient (Ccr5-/-) mice exhibit uniform lethality, compared to 30-40% mortality in wild-type controls [109]. This increased susceptibility is characterized by impaired trafficking of leukocytes—including CD4+ T cells, CD8+ T cells, natural killer (NK) cells, and macrophages—across the blood-brain barrier into the central nervous system (CNS) [109] [111]. The resulting failure to control viral replication within the brain, particularly in cortical regions including the hippocampus, leads to uncontrolled viral proliferation and fatal encephalitis [111]. Regional analysis of chemokine expression during WNV infection reveals elevated levels of CCR5 ligands (CCL4 and CCL5) in the cortices of infected CCR5-/- mice, suggesting a compensatory response that cannot overcome the trafficking defect [111].
Experimental Models and Methodologies
Murine Models of WNV Infection
Experimental Protocol:
- Animals: 8-week-old wild-type C57BL/6 mice and congenic Ccr5-/- mice (Jackson Laboratories, stock number 005427)
- Virus Inoculation: Subcutaneous footpad injection with 10² plaque-forming units (PFU) of WNV strain 3000.0259 (isolated from New York, 2000) in 50μL volume
- Clinical Monitoring: Daily assessment using standardized scoring system: 1=ruffled fur/hunched; 2=paresis/difficulty walking; 3=paralysis; 4=moribund; 5=dead
- Tissue Collection: At designated timepoints (days 6-8 post-infection), animals undergo cardiac perfusion with PBS followed by dissection of CNS regions (cortex, cerebellum, brainstem) based on anatomical boundaries
- Viral Titration: Homogenized tissues assayed by plaque formation on BHK21-15 cell monolayers
- Immune Cell Isolation: CNS leukocytes isolated via percoll gradient centrifugation and characterized by flow cytometry using antibodies to CD4, CD8β, CD11b, and CD45
- Blood-Brain Barrier Permeability: Intraperitoneal injection of sodium fluorescein (100mg/mL) with measurement of dye extravasation into CNS tissues after 45 minutes [111]
This model has demonstrated that CCR5 deficiency does not affect peripheral viral replication (in footpad, draining lymph nodes, or spleen) but specifically impairs CNS viral control, with Ccr5-/- mice exhibiting 10- to 100-fold higher viral loads in cortical regions compared to wild-type mice at day 8 post-infection [111].
Human Genetic Association Studies
Methodological Framework:
- Cohort Identification: Patients with laboratory-confirmed symptomatic WNV infection (seropositive by IgM ELISA or plaque reduction neutralization test)
- Control Populations: Healthy blood donors (pre-1999, presumed WNV-uninfected) and/or WNV-seronegative patients with similar initial symptoms
- Genotyping: DNA extraction from peripheral blood mononuclear cells using QIAamp DNA Mini Kit, PCR amplification of CCR5 locus with primers: 5'-TCATTACACCTGCAGCTCTC-3' (forward) and 5'-TGGTGAAGATAAGCCTCAC-3' (reverse)
- Product Analysis: Wild-type allele=197bp, CCR5Δ32 allele=165bp, visualized by gel electrophoresis
- Statistical Analysis: Hardy-Weinberg equilibrium testing, calculation of odds ratios with 95% confidence intervals, and survival analysis using log-rank test [110] [112]
These studies consistently demonstrate that CCR5Δ32 homozygosity increases risk for symptomatic WNV disease rather than infection acquisition, as the genotype distribution among asymptomatically infected individuals matches general population frequencies [109].
Research Reagent Solutions for CCR5 Investigation
Table 3: Essential Research Reagents for CCR5 Studies
| Reagent/Cell Line | Specific Application | Function/Utility |
|---|---|---|
| Ccr5-/- mice (B6129PF2 or C57BL/6 background) | In vivo pathogenesis studies | Determine CCR5 role in infection models through comparative analysis with wild-type controls |
| MT4CCR5 cell line | In vitro HIV infection and inhibition assays | Model system for CCR5-tropic HIV-1 infection and evaluation of antiviral strategies |
| CCR5Δ32 genotyping primers (F: 5'-TCATTACACCTGCAGCTCTC-3', R: 5'-TGGTGAAGATAAGCCTCAC-3') | Human genetic association studies | Amplify wild-type (197bp) and Δ32 (165bp) CCR5 alleles for genotype determination |
| Anti-CCR5 monoclonal antibodies (e.g., CD195) | Flow cytometry, immunohistochemistry | Detect CCR5 expression on immune cell subsets and quantify receptor density |
| CRISPR/Cas9 systems (with CCR5-specific sgRNAs) | Gene editing studies | Create CCR5-deficient cell lines or therapeutic cells for HIV resistance |
| Recombinant CCR5 ligands (CCL3, CCL4, CCL5) | Migration and signaling assays | Evaluate CCR5-mediated chemotaxis and downstream signaling events |
Therapeutic Approaches and Technological Advances
CCR5-Targeted Gene Editing Strategies
The remarkable success of CCR5Δ32/Δ32 stem cell transplantation in curing HIV-1 has spurred development of gene editing technologies to mimic this protective effect in autologous cells, thereby circumventing the need for rare matched donors and reducing transplantation morbidity [50]. Multiple platforms have emerged for creating artificial CCR5 deficiency:
Zinc Finger Nucleases (ZFNs): Among the earliest technologies applied to CCR5 editing, ZFNs employ custom-designed zinc finger proteins fused to FokI nuclease domains to induce targeted double-strand breaks in the CCR5 locus [3]. The SB-728-T clinical trial demonstrated that autologous T cells edited by ZFNs and reinfused into patients yielded acceptable safety profiles and provided virological/immunological benefits in HIV-1 infected individuals [3].
CRISPR/Cas9 Systems: The CRISPR/Cas9 platform utilizes a single guide RNA (sgRNA) to direct Cas9 nuclease to specific genomic loci, offering simplified design and high editing efficiency [53]. Clinical trial NCT03164135 assessed CRISPR/Cas9-mediated CCR5 editing in hematopoietic stem cells for patients with both HIV-1 and acute lymphoblastic leukemia, demonstrating successful engraftment and persistence of CCR5-ablated cells for over 19 months without gene-editing-related adverse events [3] [53].
Recent advances have focused on combinatorial approaches to enhance therapeutic efficacy and breadth. For instance, CRISPR/Cas9-mediated CCR5 knockout has been successfully paired with expression of C46 HIV-1 fusion inhibitor, conferring resistance to both R5-tropic and X4-tropic HIV-1 strains [53]. Similarly, multiplexed strategies now enable CCR5 disruption alongside knock-in of genes encoding potent broadly neutralizing antibodies (bNAbs) against HIV-1, creating a dual mechanism of protection through both cell-intrinsic resistance and humoral immunity [87].
Pharmacological CCR5 Antagonists
Small molecule CCR5 antagonists represent a complementary therapeutic approach, with maraviroc receiving FDA approval for HIV-1 treatment in 2007. These compounds act as allosteric inhibitors, inducing conformational changes that prevent CCR5 utilization by HIV-1 without triggering pro-inflammatory signaling [50]. While effective for viral suppression, pharmacological blockade differs fundamentally from genetic ablation in its reversibility and tissue penetration limitations, potentially mitigating flavivirus-related risks through preserved CCR5 function in some cellular compartments.
Risk Mitigation and Future Directions
The identified association between CCR5 deficiency and increased WNV susceptibility necessitates careful risk-benefit assessment in therapeutic development. Several strategies may mitigate potential adverse effects:
Geographic Considerations: CCR5-based therapies warrant cautious implementation in WNV-endemic regions, with seasonal variation in mosquito activity potentially influencing treatment timing [113]. Pre-treatment screening for WNV immunity and vaccination status could inform individual risk assessment.
Multiplexed Targeting: Combining CCR5 disruption with additional antiviral strategies could reduce dependence on CCR5 ablation alone, potentially permitting partial CCR5 function retention. Approaches include simultaneous targeting of CXCR4 to block viral tropism switching, HIV-1 fusion inhibitors to prevent viral entry regardless of coreceptor usage, and genetic elements that suppress viral replication [3] [53].
Therapeutic Monitoring: Enhanced surveillance for arboviral infections in recipients of CCR5-targeted therapies, with prompt diagnostic evaluation for flaviviruses in cases of febrile illness or neurological symptoms following treatment [112].
Future research priorities include elucidating whether CCR5 deficiency increases susceptibility to other neurotropic flaviviruses (e.g., Japanese encephalitis virus, tick-borne encephalitis virus) [109], defining threshold levels of CCR5 function required for CNS immune protection while maintaining HIV-1 resistance, and developing refined gene editing approaches that preserve specific CCR5-dependent immune functions while blocking HIV-1 coreceptor activity.
CCR5 represents both a promising therapeutic target and a cautionary example of genetic pleiotropy in human immunity. The CCR5Δ32 polymorphism demonstrates that naturally occurring genetic variation can simultaneously confer profound protection against one pathogen (HIV-1) while increasing vulnerability to another (WNV). This dichotomy necessitates careful risk-benefit analysis in therapeutic development, particularly as CCR5-targeted gene editing advances toward clinical application. Future success will likely depend on combinatorial approaches that achieve HIV-1 protection while preserving sufficient CCR5-mediated immunity against emerging and re-emerging viral threats, ultimately balancing the curative potential of CCR5 manipulation against its unintended immunological consequences.
The discovery that a 32-base-pair deletion (CCR5Δ32) in the C-C chemokine receptor 5 (CCR5) gene confers strong resistance to HIV-1 infection in homozygous individuals represents a foundational case study in modern genetics and therapeutic development [49] [20]. This natural loss-of-function mutation produces a nonfunctional receptor on the surface of immune cells, preventing R5-tropic HIV-1 viral entry and establishing CCR5 as a prime therapeutic target for AIDS intervention [3] [11]. The subsequent development of CCR5-targeted therapies—from small molecule antagonists to gene editing approaches—has positioned CCR5 research at the epicenter of critical debates surrounding the ethical and regulatory frameworks governing somatic and germline cell editing [114] [3]. This whitepaper examines the current technical landscape of CCR5-directed therapies, analyzes the ethical challenges posed by emerging gene editing technologies, and proposes structured regulatory considerations for researchers and drug development professionals working at this frontier.
Biological Basis of CCR5 and the Δ32 Mutation
CCR5 Structure and Function
CCR5 is a seven-transmembrane, G protein-coupled receptor (GPCR) primarily expressed on bone marrow-derived cells including lymphocytes, monocytes/macrophages, granulocytes, T cells, natural killer (NK) cells, and regulatory T (Treg) cells [11]. As a chemokine receptor, its primary physiological role involves orchestrating immune cell migration (chemotaxis) along chemokine gradients to sites of inflammation, thereby playing crucial roles in immune surveillance and inflammatory response [11]. Its natural ligands include CCL3 (MIP-1α), CCL4 (MIP-1β), CCL5 (RANTES), and CCL3L1, which are natural suppressors of HIV-1 infection [11].
The CCR5Δ32 Mutation and HIV Resistance
The CCR5Δ32 variant features a 32-base-pair deletion in the region encoding the second extracellular loop of the receptor, resulting in a frameshift and premature stop codon that produces a truncated, non-functional protein that fails to reach the cell surface [20] [11]. This mutation demonstrates a unique inheritance pattern and global distribution that suggests positive historical selection pressure:
Table 1: Population Genetics of CCR5Δ32
| Characteristic | Findings | Implications |
|---|---|---|
| Global Distribution | Highest frequency in Northern Europe (up to 16%); virtually absent in African, Asian, and Native American populations [20] [11] | Suggests origin after migration out of Africa; potential historical selection pressure in Europe |
| HIV Resistance | Homozygotes: Near-complete resistance to R5-tropic HIV-1 infection [34] [20]; Heterozygotes: ~50% receptor reduction, delayed AIDS progression, reduced viral loads [34] [20] | Validates CCR5 as therapeutic target; heterozygote benefit demonstrates partial receptor suppression may be therapeutic |
| Estimated Origin | 700-2,100 years before present [20] | Too recent for current HIV pandemic to have driven selection; suggests historical pathogen pressure (e.g., smallpox, plague) |
| Evolutionary Evidence | Strong linkage disequilibrium with microsatellite markers; 95% of Δ32 chromosomes carry identical flanking sequences [20] | Supports single mutational event followed by positive selection |
The protective mechanism is well-characterized: R5-tropic HIV-1 strains (predominant in early infection) require both CD4 and CCR5 coreceptors for cellular entry. Without functional CCR5 receptors on the cell surface, viral entry is effectively blocked in homozygous individuals [34] [20]. This natural resistance was dramatically demonstrated in the "Berlin" and "London" HIV patients who achieved long-term viral remission after receiving hematopoietic stem cell transplants from CCR5Δ32/Δ32 homozygous donors [3].
Figure 1: Molecular Mechanism of CCR5Δ32-mediated HIV Resistance
Technical Landscape of CCR5-Targeted Gene Editing
Gene Editing Platforms
Multiple gene editing platforms have been developed to target CCR5, each with distinct mechanisms and therapeutic characteristics:
Table 2: Comparison of Major Gene Editing Technologies for CCR5 Targeting
| Technology | Mechanism of Action | Advantages | Limitations and Challenges | Clinical Status for HIV |
|---|---|---|---|---|
| ZFNs | Custom-designed zinc finger proteins recognize specific DNA sequences and dimerize FokI nucleases to induce DNA cleavage [3] | Early clinical trial data (SB-728-T) demonstrating safety and virological/immunological benefits [3] | Complex design; higher risk of off-target effects; potential immunogenicity [3] | Phase 1/2 trials completed |
| TALENs | Transcription activator-like effector (TALE) proteins recognize specific DNA sequences, fused to FokI nucleases to cleave DNA [3] | Improved specificity over ZFNs; reduced off-target activity [3] | Technically demanding construction; large size challenges viral vector delivery [3] | Preclinical development |
| CRISPR/Cas9 | Single guide RNA (sgRNA) directs Cas9 nuclease to specific genomic loci for site-specific double-strand breaks [3] | Easy design; high editing efficiency; multiplex editing capability [3] | Off-target effects; PAM sequence dependency; potential immune response to Cas9 [3] | Early-phase clinical trials (NCT03164135) [3] |
| Base Editors | Fusion of Cas proteins with nucleotide deaminases enables precise single-nucleotide substitutions without double-strand breaks [3] | Avoids risks associated with DSBs (indels, chromosomal translocations) [3] | Potential off-target base editing; limited editing window [3] | Research phase |
Experimental Protocols for CCR5 Gene Editing
CRISPR/Cas9-Mediated CCR5 Disruption in Hematopoietic Stem/Progenitor Cells (HSPCs)
This protocol is adapted from clinical trial NCT03164135 investigating CRISPR/Cas9-edited HSPCs for patients with HIV and acute lymphoblastic leukemia [3]:
Materials:
- Human CD34+ HSPCs from mobilized peripheral blood
- CRISPR/Cas9 components: sgRNA targeting CCR5 (sequence: 5'-GACCGAATATTCATTACACCTGC-3'), Cas9 nuclease
- Delivery system: Electroporation system
- Culture media: Serum-free expansion media with cytokines (SCF, TPO, FLT3-L)
- Analysis: Flow cytometry for CCR5 surface expression, T7E1 assay for indel frequency, deep sequencing for off-target analysis
Procedure:
- Isolate and enrich CD34+ HSPCs using immunomagnetic selection
- Pre-stimulate cells in cytokine-containing media for 24 hours
- Prepare ribonucleoprotein (RNP) complex by incubating sgRNA with Cas9 protein
- Electroporate cells with RNP complex using optimized parameters
- Culture edited cells in vitro for functional assays or transplant into immunodeficient mice for engraftment studies
- Assess editing efficiency (indel %), CCR5 knockout efficiency (surface expression), and cell viability at 24-48 hours post-editing
- Evaluate multi-lineage differentiation potential in colony-forming unit assays
- Perform comprehensive off-target analysis at in silico-predicted sites with high sequence similarity
Multiplexed Gene Editing for Comprehensive HIV Resistance
To address limitations of single-target CCR5 editing, particularly viral escape via coreceptor switching to CXCR4, advanced protocols have developed multiplexed approaches:
Materials:
- Primary CD4+ T cells or HSPCs
- CRISPR/Cas9 system with multiple sgRNAs: CCR5-targeting, CXCR4-targeting, HIV LTR-targeting
- Delivery: AAV6 or lentiviral vectors for multiplex guide expression
- Analysis: NGS for on-target and off-target editing, viral challenge assays
Procedure:
- Design and validate sgRNAs with high on-target efficiency and minimal off-target risk for CCR5, CXCR4, and conserved HIV LTR regions
- Clone sgRNAs into multiplex expression vector system
- Deliver editing components to target cells via viral transduction or electroporation
- Sort successfully edited cells using fluorescence markers or magnetic selection
- Challenge edited cells with R5-tropic, X4-tropic, and dual-tropic HIV strains
- Measure viral replication (p24 ELISA), coreceptor usage, and cell viability over 14 days
- Sequence emerging viral variants to detect escape mutations
- Assess genomic integrity through karyotyping and transformation assays
Figure 2: Workflow for Multiplexed Gene Editing Strategy
Research Reagent Solutions
Table 3: Essential Research Reagents for CCR5 Gene Editing Studies
| Reagent/Category | Specific Examples | Function/Application |
|---|---|---|
| Gene Editing Platforms | CRISPR/Cas9 (SpCas9, SaCas9), ZFNs (SB-728-T), TALENs | Precision targeting of CCR5 locus [3] |
| Delivery Systems | Electroporation systems, AAV6 vectors, Lentiviral vectors, Lipid nanoparticles (LNPs) | Efficient intracellular delivery of editing components [3] |
| Cell Culture Materials | CD34+ isolation kits, Serum-free media with cytokines (SCF, TPO, FLT3-L), Extracellular matrix coatings | Maintenance and expansion of primary hematopoietic cells [3] |
| Analytical Tools | Flow cytometry antibodies (anti-CCR5, CD4), T7E1 assay, Next-generation sequencing, p24 ELISA | Assessment of editing efficiency and functional outcomes [3] |
| Animal Models | NSG mice, Humanized mouse models (BLT, NSG-Hu) | In vivo validation of edited cell engraftment and HIV protection [3] |
Ethical Considerations in Somatic vs. Germline Applications
The Somatic-Germline Distinction
The ethical and regulatory frameworks for gene editing diverge sharply between somatic and germline applications. Somatic editing targets non-reproductive cells, resulting in changes confined to the treated individual, while germline editing modifies reproductive cells or embryos, creating heritable changes that affect future generations [115]. The 2018 case of He Jiankui, who created the first gene-edited twins using CRISPR/Cas9 to disrupt CCR5 in embryos, exemplifies the ethical breaches possible in germline editing [114].
Key Ethical Concerns in CCR5 Gene Editing
Safety and Unintended Consequences
The primary safety concerns include both molecular and organismal effects:
- Off-target effects: Unwanted edits at genomic sites with sequence similarity to CCR5 sgRNAs [114] [3]
- On-target effects: Unwanted mutations at the CCR5 locus, including large deletions or chromosomal rearrangements [3]
- Pleiotropic effects: CCR5 modulates various physiological processes; its disruption may impact cognitive function, stroke recovery, and response to other pathogens [28] [20]
- Viral escape: Selective pressure from CCR5 disruption may promote emergence of CXCR4-tropic HIV strains with potentially greater pathogenicity [3]
Therapeutic Justification and Risk-Benefit Analysis
Current ethical frameworks prioritize somatic editing for established HIV infections where the benefits outweigh the risks [3] [115]. Germline editing for HIV prevention remains ethically problematic due to:
- Availability of effective alternatives (antiretroviral therapy, pre-exposure prophylaxis)
- Incomplete understanding of CCR5's pleiotropic functions
- Uncertain long-term consequences of germline modification [114] [116]
Equity and Access Considerations
Gene therapies currently cost $1-2 million per patient, creating substantial justice concerns [115]. The development of CCR5-based interventions raises specific equity issues:
- The CCR5Δ32 mutation is largely restricted to European populations, yet HIV disproportionately affects sub-Saharan Africa [34] [20]
- High-cost therapies may exacerbate global health disparities if accessible only to wealthy individuals or nations [115]
- Resource allocation questions arise between investing in gene therapies versus broader implementation of existing prevention strategies [115]
Regulatory Frameworks and Governance
Current Regulatory Landscape
Gene editing therapies face a complex regulatory environment that varies by jurisdiction but generally follows these principles:
Table 4: Regulatory Classification of Gene Editing Applications
| Application Type | Regulatory Status | Key Considerations | Examples |
|---|---|---|---|
| Somatic Cell Editing (Therapeutic) | Clinical trials permitted with appropriate oversight; some products approved [3] [115] | Risk-benefit analysis, long-term follow-up, manufacturing quality control | CRISPR-based therapies for sickle cell disease and β-thalassemia [115] |
| Germline Editing (Research) | Restricted or prohibited in many countries; limited to non-implantable embryos in some jurisdictions [114] [116] | Destruction of human embryos, informed consent of donors, oversight mechanisms | Basic research on mechanism of early human development [116] |
| Germline Editing (Clinical) | Widely prohibited; moratoria in place by many countries and international organizations [114] [115] | Heritable changes, absence of medical necessity, societal implications | He Jiankui case (widespread condemnation) [114] |
Emerging Governance Challenges
Distinguishing Therapy from Enhancement
As gene editing technologies advance, distinguishing therapeutic applications from enhancement becomes increasingly challenging yet ethically crucial. CCR5 research highlights this issue, as the same mutation that confers HIV resistance is also associated with enhanced cognitive function and memory in animal studies [20]. This dual effect creates potential for non-therapeutic use in individuals without medical need.
International Harmonization
The He Jiankui case demonstrated the consequences of regulatory disparities between countries [114]. Effective governance requires:
- International standards for safety and efficacy testing
- Harmonized definitions of prohibited applications
- Coordinated oversight of clinical trials and applications
- Transparency in research practices and outcomes
Future Directions and Recommendations
Technical Advancements
The next generation of CCR5 gene editing will likely focus on:
- Enhanced specificity: Improved Cas variants with reduced off-target effects
- Multiplexed approaches: Simultaneous targeting of CCR5, CXCR4, and viral reservoirs
- Delivery optimization: Non-viral delivery systems with improved tissue specificity
- Precision editing: Base and prime editing to create specific protective mutations without double-strand breaks [3]
Ethical and Regulatory Recommendations
For researchers and drug development professionals, we recommend:
- Prioritize somatic over germline applications until safety and ethical frameworks are established
- Implement comprehensive off-target analysis using multiple complementary methods
- Conduct long-term follow-up studies to identify potential late-onset effects
- Develop equitable access strategies early in product development
- Engage diverse stakeholders including patient communities, ethicists, and international partners
CCR5 research provides a powerful paradigm for understanding both the tremendous potential and profound challenges of gene editing technologies. The biological insights gained from studying the CCR5Δ32 mutation have catalyzed development of novel therapeutic approaches for HIV, while simultaneously highlighting the complex ethical considerations inherent in modifying human genetics. As technical capabilities advance, maintaining a responsive regulatory framework that balances scientific innovation with ethical responsibility remains paramount. Through continued interdisciplinary collaboration among researchers, clinicians, ethicists, and policymakers, we can harness these powerful technologies responsibly to address significant medical challenges while upholding fundamental ethical principles.
Conclusion
The journey of CCR5 research, from the discovery of its role as an HIV co-receptor to the exploitation of the protective Δ32 mutation, has fundamentally reshaped the landscape of HIV therapy and cure strategies. The foundational understanding of CCR5 biology has successfully been translated into innovative methodologies, including precision gene editing and synergistic immunotherapies, demonstrating tangible progress toward a functional cure. However, the path forward requires meticulously navigating significant challenges—viral escape via tropism switching, the precise safety of gene editing, and the pleiotropic nature of CCR5 in human health. Future research must prioritize the development of comprehensive multi-target approaches, robust personalized medicine frameworks, and accessible, scalable technologies. The convergence of CCR5-targeted strategies with broader immunotherapeutic interventions holds the most compelling promise for ultimately overcoming the remaining barriers to eradicating HIV.
References
- 1. The chemokine receptor CCR5: multi-faceted hook for HIV-1 [https://retrovirology.biomedcentral.com/articles/10.1186/s12977-024-00634-1]
- 2. How Mechanisms of HIV Entry Govern AIDS Pathogenesis [https://www.sciencedirect.com/science/article/abs/pii/S002228361830634X]
- 3. CCR5 gene editing and HIV immunotherapy - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC12213807/]
- 4. CCR5 [https://en.wikipedia.org/wiki/CCR5]
- 5. CCR5 Pathway in Macrophages [https://www.thermofisher.com/us/en/home/life-science/antibodies/antibodies-learning-center/antibodies-resource-library/cell-signaling-pathways/ccr5-pathway-macrophages.html]
- 6. Molecular Mechanism of HIV-1 Entry - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6744290/]
- 7. Text has enhanced contrast | ACT Rule | WAI [https://www.w3.org/WAI/standards-guidelines/act/rules/09o5cg/]
- 8. Text has enhanced contrast - Rules - Siteimprove [https://alfa.siteimprove.com/rules/sia-r66]
- 9. HIV-1 Entry and Membrane Fusion Inhibitors - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC8146413/]
- 10. CCR5: From Natural Resistance to a New Anti-HIV Strategy [https://pmc.ncbi.nlm.nih.gov/articles/PMC3185609/]
- 11. CCR5 as a Coreceptor for Human Immunodeficiency Virus ... [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2022.835994/full]
- 12. The CCR5-Delta32 Genetic Polymorphism and HIV-1 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6227735/]
- 13. The first structure of HIV-1 gp120 with CD4 and CCR5 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6317214/]
- 14. HIV-1 Env trimers asymmetrically engage CD4 receptors in ... [https://www.nature.com/articles/s41586-023-06762-6]
- 15. Conformational Changes in Cell Surface HIV-1 Envelope ... [https://www.sciencedirect.com/science/article/pii/S002192581838596X]
- 16. FUSION STAGE OF HIV-1 ENTRY DEPENDS ON VIRUS- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5558241/]
- 17. CCR5 as a Coreceptor for Human Immunodeficiency Virus ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8829453/]
- 18. CCR5Δ32 mutation and HIV infection: basis for curative ... [https://pubmed.ncbi.nlm.nih.gov/26143158/]
- 19. CCR5Δ32 Protein Expression and Stability Are Critical for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1951285/]
- 20. CCR5-Δ32 [https://en.wikipedia.org/wiki/CCR5-%CE%9432]
- 21. CCR5 gene editing and HIV immunotherapy [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2025.1590690/full]
- 22. High frequency CCR5 editing in human hematopoietic ... [https://www.nature.com/articles/s41467-025-55873-3]
- 23. CCR5-Δ32 biology, gene editing, and warnings for the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7106751/]
- 24. The evolutionary history of the CCR5-Δ32 HIV-resistance ... [https://www.sciencedirect.com/science/article/pii/S1286457904003636]
- 25. Beyond HIV infection: Neglected and varied impacts of CCR5 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7260533/]
- 26. SUSTAINED HIV-1 REMISSION FOLLOWING ... [https://www.natap.org/2019/CROI/croi_32.htm]
- 27. Genetic Polymorphisms in the Open Reading Frame of ... [https://www.nature.com/articles/s41598-019-44136-z]
- 28. Beyond HIV infection: Neglected and varied impacts of ... [https://www.sciencedirect.com/science/article/pii/S0168170220302938]
- 29. Distribution of CCR5-Delta32, CCR2-64I, and SDF1-3'A ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12102844/]
- 30. Distribution of CCR5-Δ32 and HLA-B*57:01 alleles in HIV ... [https://www.nature.com/articles/s41439-025-00321-3]
- 31. The CCR5Δ32 allele as an HIV infection resistance marker [https://www.sciencedirect.com/science/article/pii/S2452014423001188?dgcid=rss_sd_all&]
- 32. The Geographic Spread of the CCR5 Δ32 HIV-Resistance Allele [https://pmc.ncbi.nlm.nih.gov/articles/PMC1255740/]
- 33. Frequencies of gene variant CCR5-Δ32 in 87 countries ... [https://www.sciencedirect.com/science/article/pii/S0198885917305104]
- 34. The coreceptor mutation CCR5Δ32 influences ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC56941/]
- 35. CCR5 inhibitors: Emerging promising HIV therapeutic strategy [https://pmc.ncbi.nlm.nih.gov/articles/PMC3168031/]
- 36. In-depth virological and immunological characterization of HIV ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10033413/]
- 37. Detection of CCR5Δ32 Mutant Alleles in Heterogeneous ... [https://www.frontiersin.org/journals/molecular-biosciences/articles/10.3389/fmolb.2022.805931/full]
- 38. The Role of the Chemokine Receptor Gene CCR5 and Its ... [https://wwwnc.cdc.gov/eid/article/3/3/97-0302_article]
- 39. CC Chemokine Receptor 5 Genotype and Susceptibility to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3319124/]
- 40. CCR5 promoter activity correlates with HIV disease ... [https://www.nature.com/articles/s41598-017-00192-x]
- 41. Homozygous and heterozygous CCR5-Delta32 genotypes are ... [https://pubmed.ncbi.nlm.nih.gov/11511825/]
- 42. Concordance of CCR5 Genotypes that Influence Cell ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3071050/]
- 43. Progress in structure-based drug development targeting ... [https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2025.1603950/full]
- 44. Clinical use of CCR5 inhibitors in HIV and beyond [https://translational-medicine.biomedcentral.com/articles/10.1186/1479-5876-9-S1-S9]
- 45. Profile of maraviroc: a CCR5 antagonist in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3218686/]
- 46. Maraviroc: a new CCR5 antagonist [https://pubmed.ncbi.nlm.nih.gov/19622053/]
- 47. Progress in structure-based drug development targeting ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12183272/]
- 48. Aphios Nanotherapy Targets CCR5 for Alzheimer's [https://aphios.com/about/press-releases/aphios-nanotherapy-alzheimers-ccr5-interview-2025/]
- 49. Legacy of a magic gene—CCR5-∆32: From discovery to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10962972/]
- 50. CCR5Δ32 mutation and HIV infection: basis for curative ... [https://www.sciencedirect.com/science/article/abs/pii/S1879625715000966]
- 51. CRISPR-Cas9, TALENs and ZFNs - the battle in gene editing [https://www.ptglab.com/news/blog/crispr-cas9-talens-and-zfns-the-battle-in-gene-editing/?srsltid=AfmBOopVVOM_t31GAtP8knCoAJ4gEmJsYMS2vQ5CIsOCNR1X6A8qlJfo]
- 52. Comparing Gene Editing Platforms: CRISPR vs. Traditional ... [https://blog.crownbio.com/comparing-gene-editing-platforms-crispr-vs-traditional-methods]
- 53. CRISPR/Cas9 genome editing of CCR5 combined with ... [https://www.nature.com/articles/s41598-024-61626-x]
- 54. The comparison of ZFNs, TALENs, and SpCas9 by GUIDE ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8655392/]
- 55. HIV-1 remission following CCR5Δ32/Δ32 haematopoietic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7275870/]
- 56. HIV-1 remission following CCR5Δ32/Δ32 haematopoietic ... [https://www.nature.com/articles/s41586-019-1027-4]
- 57. The next Berlin Patient: Another man cured of HIV after ... [https://www.eatg.org/hiv-news/the-next-berlin-patient-another-man-cured-of-hiv-after-stem-cell-transplant/]
- 58. Evidence for HIV-1 cure after CCR5Δ32/Δ32 allogeneic ... [https://pubmed.ncbi.nlm.nih.gov/32169158/]
- 59. Second Berlin patient has unusual immune response that ... [https://www.aidsmap.com/news/oct-2025/second-berlin-patient-has-unusual-immune-response-seems-have-removed-his-hiv]
- 60. Multiplexed Guide RNA Expression Leads to Increased ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10443033/]
- 61. Recent Advances targeting CCR5 for Cancer and its Role ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6810651/]
- 62. Mechanisms, combination therapy, and biomarkers in cancer ... [https://biosignaling.biomedcentral.com/articles/10.1186/s12964-024-01711-w]
- 63. CCR5-edited CD4+ T cells augment HIV-specific immunity ... [https://www.jci.org/articles/view/144486]
- 64. Advancing gene editing therapeutics: Clinical trials and ... [https://www.sciencedirect.com/science/article/pii/S2162253125002203]
- 65. CAR-T cell therapy for cancer: current challenges and ... [https://www.nature.com/articles/s41392-025-02269-w]
- 66. Modeling Unique Patients in Humanized Mice [https://pmc.ncbi.nlm.nih.gov/articles/PMC5835123/]
- 67. Transgenic mouse expressing human CCR5 as a model ... [https://www.sciencedirect.com/science/article/abs/pii/S0014299905006163]
- 68. The biology of CCR5 and CXCR4 - PMC - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC2718543/]
- 69. How HIV changes its tropism: evolution and adaptation? - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC2697388/]
- 70. CCR5/CXCR4 Dual Antagonism for the Improvement of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6384722/]
- 71. Immune activation correlates with and predicts CXCR4 co ... [https://www.nature.com/articles/s41598-020-71699-z]
- 72. HIV coreceptor tropism determination and mutational ... [https://www.nature.com/articles/srep21280]
- 73. Modelling binding between CCR5 and CXCR4 receptors and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3833284/]
- 74. Discovery of HIV entry inhibitors via a hybrid CXCR4 and ... [https://www.sciencedirect.com/science/article/abs/pii/S0928098720303250]
- 75. In-depth virological and immunological characterization of ... [https://www.nature.com/articles/s41591-023-02213-x]
- 76. Closing the Door with CRISPR: Genome Editing of CCR5 and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9326690/]
- 77. Off-target effects in CRISPR/Cas9 gene editing - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10034092/]
- 78. Off-target Effects in CRISPR/Cas9-mediated Genome ... [https://www.sciencedirect.com/science/article/pii/S216225311630049X]
- 79. CRISPR Off-Target Effects: Mechanisms and Solutions [https://lifesciences.danaher.com/us/en/library/crispr-off-target-effects.html]
- 80. CCR5 and Biological Complexity: The Need for Data ... [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2021.790041/full]
- 81. Co-receptor signaling in the pathogenesis of neuroHIV [https://retrovirology.biomedcentral.com/articles/10.1186/s12977-021-00569-x]
- 82. Frequencies of 32 base pair deletion of the (Δ32) allele of ... [https://www.sciencedirect.com/science/article/abs/pii/S1567134802000278]
- 83. Profound phenotypic and epigenetic heterogeneity of the ... [https://www.nature.com/articles/s41590-022-01371-3]
- 84. Heterogeneity of HIV-1 latent reservoirs - PMC - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC10631589/]
- 85. Detection of CCR5Δ32 Mutant Alleles in Heterogeneous ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8898955/]
- 86. In pursuit of an HIV cure: from stem cell transplants to gene ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12446371/]
- 87. Multilayered HIV-1 resistance in HSPCs through CCR5 ... [https://www.nature.com/articles/s41467-025-58371-8]
- 88. Sustained HIV remission after allogeneic hematopoietic ... [https://www.nature.com/articles/s41591-024-03277-z]
- 89. Targeting CCR5 as a Component of an HIV-1 Therapeutic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8811197/]
- 90. The global distribution of CCR5 delta 32 polymorphism [https://pmc.ncbi.nlm.nih.gov/articles/PMC3344725/]
- 91. Automated production of CCR5-negative CD4 + -T cells in ... [https://www.nature.com/articles/s41434-021-00259-5]
- 92. Efficient manufacturing and engraftment of CCR5 gene- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10415663/]
- 93. 2025 cell and gene challenges: Scalability, supply chain ... [https://www.cellgenetherapyreview.com/3975-Featured-Articles/617533-2025-cell-and-gene-challenges-Scalability-supply-chain-and-manufacturing/]
- 94. Targeting CCR5 as a Component of an HIV-1 Therapeutic ... [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2021.816515/full]
- 95. CCR5 Inhibitors and HIV-1 Infection [https://www.scientificarchives.com/article/ccr5-inhibitors-and-hiv1-infection]
- 96. Patterns of inflammation and immune activation by ... [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2025.1632287/full]
- 97. Next-generation sequencing of CCR5, CXCR4, and ... [https://www.nature.com/articles/s41598-025-11843-9]
- 98. CCR5-Δ32 biology, gene editing, and warnings for the future of CRISPR-Cas9 as a human and humane gene editing tool | Cell & Bioscience | Full Text [https://cellandbioscience.biomedcentral.com/articles/10.1186/s13578-020-00410-6]
- 99. Zinc finger nuclease-mediated gene editing in ... [https://www.nature.com/articles/s41598-024-74716-7]
- 100. CRISPR-Cas9-mediated gene disruption of HIV-1 co-receptors confers broad resistance to infection in human T cells and humanized mice - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC8847835/]
- 101. Characterizing and controlling CRISPR repair outcomes in ... [https://www.nature.com/articles/s41467-025-66058-3]
- 102. Editing the human genome: where ART and science intersect [https://pmc.ncbi.nlm.nih.gov/articles/PMC6086805/]
- 103. The CCR5Δ32 allele as an HIV infection resistance marker [https://www.sciencedirect.com/science/article/abs/pii/S2452014423001188]
- 104. of CCR5-null phenotypes in humans and in... Parallel evolution [https://pubmed.ncbi.nlm.nih.gov/9707408/]
- 105. SIV Coreceptor Specificity in Natural and Non-Natural Host ... [https://pubmed.ncbi.nlm.nih.gov/29173179/]
- 106. Population size changes and selection drive... | Nature Communications [https://www.nature.com/articles/s41467-018-03990-7?error=cookies_not_supported&code=209557e8-c157-4528-953c-1ff373bc753f]
- 107. Molecular Mechanism of Action for Allosteric Modulators ... [https://www.sciencedirect.com/science/article/pii/S0021925820344367]
- 108. Modeling the allosteric modulation of CCR5 function ... [https://pubmed.ncbi.nlm.nih.gov/24050281/]
- 109. A Pocket Guide to CCR5—Neurotropic Flavivirus Edition [https://www.mdpi.com/1999-4915/16/1/28]
- 110. CCR5 deficiency increases risk of symptomatic West Nile ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2118086/]
- 111. CCR5 limits cortical viral loads during West Nile virus infection ... [https://jneuroinflammation.biomedcentral.com/articles/10.1186/s12974-015-0447-9]
- 112. Chemokine Receptor 5 Δ32 Allele in Patients with Severe ... [https://wwwnc.cdc.gov/eid/article/16/10/10-0108_article]
- 113. CCR5 thwarts West Nile virus - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC2118082/]
- 114. Ethical and scientific issues of gene-edited twin by ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7294836/]
- 115. CRISPR & Ethics [https://innovativegenomics.org/crisprpedia/crispr-ethics/]
- 116. Ethical issues related to research on genome editing in ... [https://www.sciencedirect.com/science/article/pii/S2001037019305173]